2222
Federated training of deep learning models for prostate cancer segmentation on MRI: A simulation study1Georgia Institute of Technology, Atlanta, GA, United States, 2University of Pennsylvania, Philadelphia, PA, United States, 3Rhino Health, Boston, MA, United States, 4Emory University, Atlanta, GA, United States, 5Atlanta VA Medical Center, Atlanta, GA, United States
Synopsis
Keywords: Diagnosis/Prediction, Segmentation
Motivation: Site and scanner specific variations in prostate MRI impact performance of deep learning (DL) based models. Federated learning allows for privacy preserving training of DL models without the need for data sharing.
Goal(s): In this study, we train DL models for prostate cancer segmentation on MRI using the Rhino Health federated computing platform.
Approach: We adopt 3D UNet architecture to train the DL models on 2 publicly available datasets.
Results: DL models trained using a federated approach result in more generalizable models compared to those trained on single site data.
Impact: Successful development of deep learning based prostate cancer segmentation models on MRI using federated learning will result in reproducible and generalizable models. These can enhance clinical adoption and potentially improve downstream diagnostic and treatment workflows for prostate cancer.
Introduction
Automated segmentation of prostate cancer (PCa) regions of interest (ROI) on MRI offers several advantages for downstream PCa diagnosis and radiotherapy planning. Deep learning (DL) has been extensively shown[1], [2] to achieve very good performance for PCa segmentation, however, DL models are susceptible to batch artifacts arising from site and scanner specific variations[3], impacting reliability and clinical translation. Access to multi-site datasets is becoming increasingly challenging due to patient privacy concerns, data storage and transfer costs. Federated learning (FL) allows for training DL models without the need for data sharing outside institutional firewall, preserving data privacy[4]. In this study, we leverage Rhino Health federated computing platform (FCP)[5] to simulate federated training of DL for PCa segmentation on MRI, using publicly available datasets. We compare performance of federated and single site training of DL models.Methods
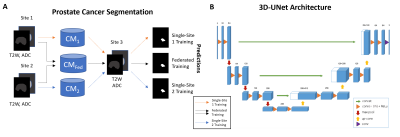
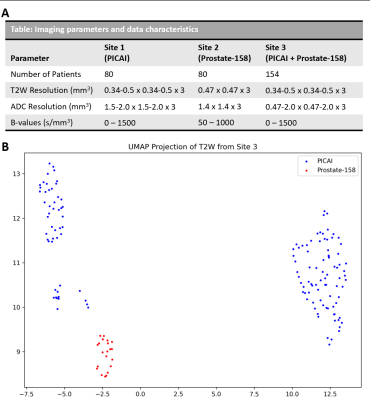
In this study, we leveraged N=314 studies from 2 publicly available datasets (PICAI challenge[6] N=213 and Prostate-158[7] N=101) consisting of 3T multi-parametric prostate MRI scans. Studies that contained expert radiologist drawn PCa annotations were selected from the original datasets. Gleason Grade Group (GGG) from targeted biopsy for these patients were used to determine clinically significant PCa (csPCa) defined as GGG>1. Patients without csPCa were excluded from the study. Two 3D U-Net[8] based DL models from MONAI open source framework were defined, one each for prostate cancer segmentation. Dice loss with an Adam optimizer was used, and learning rate=2e-3, batch size=3 were set as hyperparameters. Biparametric MRI (T2-weighted (T2W) MRI and apparent diffusion coefficient (ADC) maps[9]) were used for training the models. Datasets were partitioned into 3 sites simulated for federated learning: S1(PICAI; N=80), S2(P158; N=80) and S3(N=154; 133 PICAI and 21 P158). Models CM1, CM2 and aggregate model CMFed were trained on S1, S2 and S1+S2 respectively for prostate cancer segmentation (Fig 1) using the Rhino FCP platform[5]. Models were trained for N=100 epochs (aggregated over 4 rounds for CMFed) employing an early stopping criteria. All the models were validated on S3 and evaluated for performance in terms of dice similarity coefficient (DSC) and Jaccard Index (JI). DSC evaluates intersection over disjoint union while JI computes the intersection over union between predicted ROI and the ground truth ROI.Results and Discussion
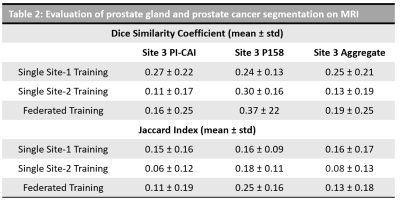
The dataset characteristics and imaging parameters are presented in Fig 1. We observed significant differences in the intensities across datasets from PICAI and Prostate-158 datasets used in S3 as indicated in the UMAP in Fig 1. Models CM1, CM2 resulted in a DSC= 0.25 ± 0.21 and 0.13 ± 0.19, JI= 0.16 ± 0.17 and 0.08 ± 0.13 on S3 respectively. The aggregate model CMFed resulted in DSC=0.19 ± 0.25 and JI=0.13 ± 0.18 on S3. When we computed the metrics on PICAI and Prostate-158 patients separately within S3, we observed that CM1 performed on studies from PICAI, and CM2 performed on studies from Prostate-158, since they belonged the same dataset. The aggregate model trained using federated learning approach, resulted in improved performance on cross institutional datasets DSC = 0.37 vs 0.24, 0.16 vs 0.11, as shown in Table 2.Discussion
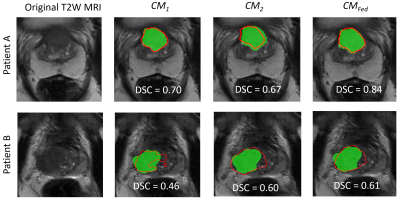
We observe that single site model are susceptible to site and scanner specific variations and federated learning results in more generalizable models (Fig 3). When we performed a subset analysis on patients within S3, we observed that single site models performed well on respective datasets while the segmentation performance was low on cross institution dataset. While the overall dice coefficients are relatively low, we observe trends that show federated learning results in more generalizable and reliable segmentations than single site training. Previous studies[10], [11] have explored federated learning for prostate MRI, however, they primarily focused on the task of prostate gland segmentation. In our study, we focused on the more challenging task of prostate cancer segmentation. Additionally, we demonstrated feasibility using commercial federated learning platform (Rhino FCP) and performed simulated federated learning. Nevertheless, there is no practical difference since the process is essentially the same. The performance metrics reported in our study are relatively low since we used a basic UNet framework and with use of state of the art segmentation models such as SAM[12], vision transformers[13], the segmentation performance will be significantly higher.Conclusion
We demonstrated that federated learning results in more generalizable deep learning model for prostate cancer segmentation on MRI, robust to site and scanner specific variations, compared to models trained on single site. Future work involves validation on large institutional, datasets and leverage state of the art deep learning models for improved segmentation performance.Acknowledgements
Research reported in this publication was supported by the National Cancer Institute under award numbers R01CA268287A1, U01CA269181, R01CA26820701A1, R01CA249992-01A1, R01CA202752-01A1, R01CA208236-01A1, R01CA216579-01A1, R01CA220581-01A1, R01CA257612-01A1, 1U01CA239055-01, 1U01CA248226-01, 1U54CA254566-01, National Heart, Lung and Blood Institute 1R01HL15127701A1, R01HL15807101A1, U01CA113913, National Institute of Biomedical Imaging and Bioengineering 1R43EB028736-01, VA Merit Review Award IBX004121A from the United States Department of Veterans Affairs Biomedical Laboratory Research and Development Service, the Office of the Assistant Secretary of Defense for Health Affairs, through the Breast Cancer Research Program (W81XWH-19-1-0668), the Prostate Cancer Research Program (W81XWH-20-1-0851), the Lung Cancer Research Program (W81XWH-18-1-0440 and W81XWH-20-1-0595), the Peer Reviewed Cancer Research Program (W81XWH-18-1-0404, W81XWH-21-1-0345, and W81XWH-21-1-0160), (W81XWH-22-1-0236) the Kidney Precision Medicine Project (KPMP) Glue Grant, sponsored research agreements from Bristol Myers-Squibb, Boehringer-Ingelheim, Eli-Lilly, and AstraZeneca, American Cancer Society Institutional Research Grant from Winship Cancer Institute, and Winship Invest$ Pilot Grant from the Winship Cancer Institute.References
[1] A. Hiremath et al., “An integrated nomogram combining deep learning, Prostate Imaging–Reporting and Data System (PI-RADS) scoring, and clinical variables for identification of clinically significant prostate cancer on biparametric MRI: a retrospective multicentre study,” Lancet Digit. Health, vol. 3, no. 7, pp. e445–e454, Jul. 2021, doi: 10.1016/S2589-7500(21)00082-0.
[2] B. Turkbey and M. A. Haider, “Deep learning-based artificial intelligence applications in prostate MRI: brief summary,” Br. J. Radiol., vol. 95, no. 1131, p. 20210563, Mar. 2022, doi: 10.1259/bjr.20210563.
[3] R. Kushol, P. Parnianpour, A. H. Wilman, S. Kalra, and Y.-H. Yang, “Effects of MRI scanner manufacturers in classification tasks with deep learning models,” Sci. Rep., vol. 13, no. 1, p. 16791, Oct. 2023, doi: 10.1038/s41598-023-43715-5. [4] N. Rieke et al., “The future of digital health with federated learning,” NPJ Digit. Med., vol. 3, p. 119, 2020, doi: 10.1038/s41746-020-00323-1.
[5] I. Dayan et al., “Federated learning for predicting clinical outcomes in patients with COVID-19,” Nat. Med., vol. 27, no. 10, pp. 1735–1743, Oct. 2021, doi: 10.1038/s41591-021-01506-3.
[6] A. Saha et al., “Artificial Intelligence and Radiologists at Prostate Cancer Detection in MRI: The PI-CAI Challenge (Study Protocol),” Zenodo, Jun. 2022. doi: 10.5281/ZENODO.6667655.
[7] L. C. Adams et al., “Prostate158 - An expert-annotated 3T MRI dataset and algorithm for prostate cancer detection,” Comput. Biol. Med., vol. 148, p. 105817, Sep. 2022, doi: 10.1016/j.compbiomed.2022.105817.
[8] M. J. Cardoso et al., “MONAI: An open-source framework for deep learning in healthcare,” 2022, doi: 10.48550/ARXIV.2211.02701.
[9] J. Cho et al., “Biparametric versus multiparametric magnetic resonance imaging of the prostate: detection of clinically significant cancer in a perfect match group,” Prostate Int., Feb. 2020, doi: 10.1016/j.prnil.2019.12.004.
[10] H. R. Roth et al., “Federated Whole Prostate Segmentation in MRI with Personalized Neural Architectures,” in Medical Image Computing and Computer Assisted Intervention – MICCAI 2021, vol. 12903, M. De Bruijne, P. C. Cattin, S. Cotin, N. Padoy, S. Speidel, Y. Zheng, and C. Essert, Eds., in Lecture Notes in Computer Science, vol. 12903. , Cham: Springer International Publishing, 2021, pp. 357–366. doi: 10.1007/978-3-030-87199-4_34.
[11] K. V. Sarma et al., “Federated learning improves site performance in multicenter deep learning without data sharing,” J. Am. Med. Inform. Assoc. JAMIA, vol. 28, no. 6, pp. 1259–1264, Jun. 2021, doi: 10.1093/jamia/ocaa341.
[12] J. Ma, Y. He, F. Li, L. Han, C. You, and B. Wang, “Segment Anything in Medical Images,” 2023, doi: 10.48550/ARXIV.2304.12306.
[13] Z. Liu et al., “Swin Transformer: Hierarchical Vision Transformer using Shifted Windows,” 2021, doi: 10.48550/ARXIV.2103.14030.
Figures