2221
Federated image-to-image MRI translation from heterogeneous multiple-sites data1Sano Centre for Computational Medicine, Kraków, Poland, 2AGH University of Science and Technology, Kraków, Poland, 3Laboratorio de Procesado de Imagen (LPI), ETSI Telecomunicación, Universidad de Valladolid, Valladolid, Spain
Synopsis
Keywords: AI/ML Image Reconstruction, Machine Learning/Artificial Intelligence, Federated Learning, Image-to-image Translation
Motivation: Applying machine learning (ML) in MRI necessitates the development of large and diverse datasets, which is a challenging process. Federated learning (FL) is a new frontier in ML that offers the possibility of multi-site data aggregation.
Goal(s): In our study, we examine a traditional deep convolutional neural network applied to multiple sources with that of the FL technique using different aggregation methods.
Approach: As a proof-of-concept, we employ four publicly available MRI datasets and carry out image-to-image translation between T1- and T2-weighted scans.
Results: Our findings suggest that the FL generalizes the model more effectively than using models trained at each site separately.
Impact: Our research demonstrated the crucial role of federated learning in medical imaging. It also emphasized the significance of selecting an appropriate aggregation algorithm considering the data type and degree of heterogeneity.
Introduction
Data synthesis in medical imaging is a new frontier in MRI aimed at reducing the acquisition time or filling in missing data in multi-parameter acquisitions1,2,3. It can also limit the patient's exposure to harmful factors4,5 or help obtain quantitative maps based on T1- and T2-weighted data4. Clinical applications of data synthesis include its use in neuro-oncology6, particularly in brain tumour classification6 and radiotherapy planning7. Integrating data from multiple centres with varying acquisition protocols is essential for utilizing deep learning models in image translation8,9. However, this process raises concerns about maintaining the privacy of sensitive medical data and the limited ability of neural networks to generalize. Federated learning (FL) is a promising approach to address both issues10,11. It involves aggregating information about data from various centres to train a shared model without transferring the data between the sources.Methods and materials:
Data synthesis: The image-to-image translation approach employed the U-Net model12, featuring an initial block with 64 channels, followed by a progressive doubling of channels until reaching the bottleneck layer with 1024 channels. The decoder utilized a reverse architecture. The training was conducted with Adam13 optimizer, initial learning rate of 0.001, and batch size 32. Previously utilzed loss for identical task14 was improved by summing MSE with DSSIM. For single-dataset training (no FL applied), the number of epochs was set to 50, which was sufficient for convergence.Federated learning: All the aggregation methods shared the same FL parameters: 4 local epochs, 32 global epochs (rounds), and 1.0 fraction fit, meaning— all the clients were included in every iteration:
- FedAvg15 : a weighted average of the model parameters;
- FedAdam16: an adaptive optimization method; parameters used in the study: $$$\tau=0.001$$$, $$$\eta=0.1$$$, $$$\eta_l=0.1$$$, $$$\beta_1=0.9$$$, $$$\beta_2=0.99$$$;
- FedBN17: weighted average excluding normalization layers;
- FedCostWAvg18: considers the loss change in computing the weighted average; parameters used in the study: $$$\alpha=0.5$$$;
- FedMRI19 model divided into the global encoder and local decoders. The utilized version is referred to as FedMRI$$$\dagger$$$ in the paper19 (without the extra loss component).
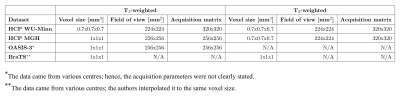
Datasets: We use the following datasets (see Fig. 1):
- HCP WU-Minn20: 104 healthy subjects, aged 22-35y;
- HCP MGH21: 26 healthy subjects, aged 20-59y;
- OASIS-322: 125 subjects, some at various stages of cognitive decline, aged 42-95y;
- BraTS23,24,25: further divided into low and high-grade gliomas datasets (LGG, HGG) of 76 and 50 subjects respectively.
Data preprocessing: The raw data were skull stripped (FSL, bet26) and co-registered between T1- and T2-weighted volumes (FSL, flirt27 and fnirt28). Voxel values were normalized to the 0-1 range, and only slices containing a significant portion of the brain image were selected for further analysis.
Results
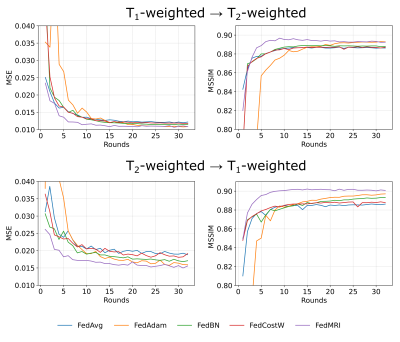
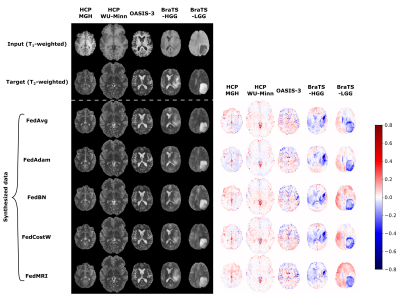
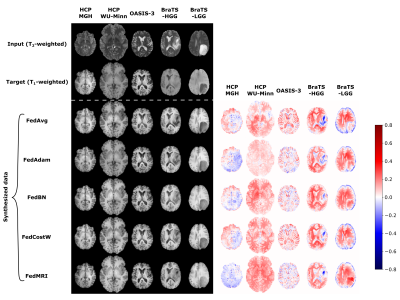
In Fig. 2, we present the averaged mean square error (MSE) and mean structural similarity index measure (MSSIM) values among all clients as a function of global rounds. The selection of an FL-based algorithm affects not only the correctness of the results obtained but also the speed and stability of model training. Among the various methods used, the best results are achieved by the FedMRI aggregation.Next, in Fig. 3 and Fig. 4, we visually inspect the translation between T1- and T2-weighted MRI data, respectively, for randomly selected subjects from all datasets in the axial slice. This experiment exposes different limitations associated with each of the aggregation methods, especially in the glioma area, and confirms that FedMRI in a broad experimental setting is the most effective method.
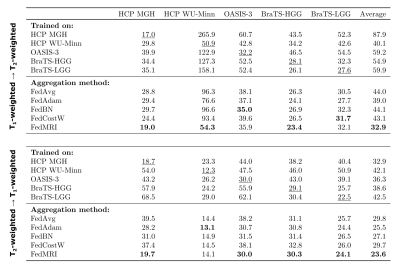
In Fig. 5, we expand upon previous research by testing the performance of a model trained on a single dataset when applied to all clients' data and tabulating the relative error values. These analyses again demonstrate that the FedMRI technique yields the best results. Our findings confirm the critical role of FL, i.e. the simple aggregation algorithm (FedAvg) yields a model performing generally more accurately than the traditionally trained.
Discussion and conclusion
In this study, we evaluated the effectiveness of different federated learning algorithms for image-to-image T1- and T2-weighted translation. We compared the performance outcomes of a standard ML approach with the training results in the FL setting. Implementing FL not only enhances the security of medical data but also improves the performace of the acquired model. However, using the basic aggregation methods is suboptimal for MRI data due to high heterogeneity arising from variations in scanner models and acquisition protocols. To achieve better results, it is necessary to employ more advanced learning techniques, such as FedMRI, specifically designed for this type of data.Acknowledgements
Jan Fiszer and Dominika Ciupek contributed equally. The numerical experiment was possible through computing allocation on the Ares and Athena systems at ACC Cyfronet AGH under the grant PLG/2023/016117. Tomasz Pieciak acknowledges the Polish National Agency for Academic Exchange for grant PPN/BEK/2019/1/00421 under the Bekker programme and the Ministry of Science and Higher Education (Poland) under the scholarship for outstanding young scientists (692/STY/13/2018). This work is supported by the European Union’s Horizon 2020 research and innovation programme under grant agreement Sano No 857533 and the International Research Agendas Programme of the Foundation for Polish Science No MAB PLUS/2019/13.Data were provided in part by OASIS, OASIS-3: Longitudinal Multimodal Neuroimaging: Principal Investigators: T. Benzinger, D. Marcus, J. Morris; NIH P30 AG066444, P50 AG00561, P30 NS09857781, P01 AG026276, P01 AG003991, R01 AG043434, UL1 TR000448, R01 EB009352. AV-45 doses were provided by Avid Radiopharmaceuticals, a wholly owned subsidiary of Eli Lilly.Data collection and sharing for this project was provided by the Human Connectome Project (HCP; Principal Investigators: Bruce Rosen, M.D., Ph.D., Arthur W. Toga, Ph.D., Van J. Weeden, MD). HCP funding was provided by the National Institute of Dental and Craniofacial Research (NIDCR), the National Institute of Mental Health (NIMH), and the National Institute of Neurological Disorders and Stroke (NINDS). HCP data are disseminated by the Laboratory of Neuro Imaging at the University of Southern California.References
- Yang Q., Li N., Zhao Z., et al. MRI cross-modality image-to-image translation. Scientific Reports. 2020;10(1):3753.
- Zhou T., Fu H., Chen G., et al. Hi-net: hybrid-fusion network for multi-modal MR image synthesis. IEEE Transactions on Medical Imaging. 2020;39(9):2772-2781.
- Moshe Y. H., Buchsweiler Y., Teicher M., Artzi M. Handling missing MRI data in brain tumors classification tasks: Usage of synthetic images vs. duplicate images and empty images. Journal of Magnetic Resonance Imaging. 2023.
- Armanious K., Jiang C., Fischer M., et al. MedGAN: Medical image translation using GANs. Computerized Medical Imaging and Graphics. 2020;79:101684.
- Moya-Sáez E., Peña-Nogales Ó., de Luis-García R., Alberola-López C. A deep learning approach for synthetic MRI based on two routine sequences and training with synthetic data. Computer Methods and Programs in Biomedicine. 2021;210:106371.
- Brou Boni K. N., Klein J., Gulyban A., et al. Improving generalization in MR‐to‐CT synthesis in radiotherapy by using an augmented cycle generative adversarial network with unpaired data. Medical Physics. 2021;48(6):3003-3010.
- Moya-Sáez E., de Luis-García R., Alberola-López C. Toward deep learning replacement of gadolinium in neuro-oncology: A review of contrast-enhanced synthetic MRI. Frontiers in Neuroimaging. 2023;2:1055463.
- Iglesias J. E., Billot B., Balbastre Y., et al. SynthSR: A public AI tool to turn heterogeneous clinical brain scans into high-resolution T1-weighted images for 3D morphometry. Science Advances. 2023;9(5):eadd3607.
- Billot B., Magdamo C., Cheng Y., et al. Robust machine learning segmentation for large-scale analysis of heterogeneous clinical brain MRI datasets. Proceedings of the National Academy of Sciences. 2023;120(9):e2216399120.
- Rieke N., Hancox J., Li W., et al. The future of digital health with federated learning. NPJ Digital Medicine. 2020;3(1):119.
- Li T., Sahu A. K., Talwalkar A., Smith V. Federated learning: Challenges, methods, and future directions. IEEE Signal Processing Magazine. 2020;37(3):50-60.
- Ronneberger O., Fischer P., Brox T. U-net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference. Springer International Publishing. 2015;234-241.
- Kingma D. P., Ba J. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980. 2014.
- Osman A. F., Tamam N. M. Deep learning‐based convolutional neural network for intramodality brain MRI synthesis. Journal of Applied Clinical Medical Physics. 2022;23(4):e13530.
- McMahan B., Moore E., Ramage D., et al. Communication-efficient learning of deep networks from decentralized data. In Artificial Intelligence and Statistics. PMLR. 2017;1273-1282.
- Reddi S., Charles Z., Zaheer M., et al. Adaptive federated optimization. arXiv preprint arXiv:2003.00295. 2020.
- Li X., Jiang M., Zhang X., et al. FedBn: Federated learning on non-iid features via local batch normalization. arXiv preprint arXiv:2102.07623. 2021.
- Mächler L., Ezhov I., Kofler F., et al. FedCostWAvg: A new averaging for better Federated Learning. In International MICCAI Brainlesion Workshop. Cham: Springer International Publishing. 2021;383-391.
- Feng C. M., Yan Y., Wang S., et al. Specificity-preserving federated learning for MR image reconstruction. IEEE Transactions on Medical Imaging, 2023;42(7):2010-221.
- Menze B. H., Jakab A., Bauer S., et al. The Multimodal Brain Tumor Image Segmentation Benchmark (BRATS). IEEE Transactions on Medical Imaging. 2015;34(10):1993-2024.
- Bakas S., Akbari H., Sotiras A., et al. Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features. Nature Scientific Data. 2017;4:170117.
- Bakas S., Reyes M., Jakab A., et al. Identifying the Best Machine Learning Algorithms for Brain Tumor Segmentation, Progression Assessment, and Overall Survival Prediction in the BRATS Challenge. arXiv preprint arXiv:1811.02629. 2018.
- Fan Q., Witzel T., Nummenmaa A., et al. MGH-USC Human Connectome Project datasets with ultra-high b-value diffusion MRI. NeuroImage. 2015;124:1108-1114.
- LaMontagne P. J., Benzinger T. L., Morris J. C., et al. OASIS-3: longitudinal neuroimaging, clinical, and cognitive dataset for normal aging and Alzheimer disease. MedRxiv. 2019;2019-12.
- Van Essen D. C., Smith S. M., Barch D. M., et al. The WU-Minn human connectome project: an overview. NeuroImage. 2013;80:62-79.
- Jenkinson M., Pechaud M., Smith S. BET2: MR-based estimation of brain, skull and scalp surfaces. In Eleventh annual meeting of the organization for human brain mapping. 2005;17(3):167.
- Jenkinson M., Bannister P., Bradyv M., Smith S. Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage. 2002;17(2):825-841.
- Andersson J. L., Jenkinson M., Smith S. Non-linear registration, aka Spatial normalisation. FMRIB technical report TR07JA2. FMRIB Analysis Group of the University of Oxford. 2007;2(1):e21.
Figures