2219
Multi-domain and Uni-domain Fusion for domain-generalizable fMRI-based phenotypic prediction1National University of Singapore, Singapore, Singapore, 2Heinrich-Heine University Düsseldorf, Düsseldorf, Germany, 3Yale University, New Haven, CT, United States, 4McGill University, Montreal, QC, Canada, 5Rutgers University, Piscataway, NJ, United States
Synopsis
Keywords: Diagnosis/Prediction, fMRI (resting state), functional connectivity, phenotypic prediction, meta-learning, transfer learning
Motivation: Resting-state functional connectivity (RSFC) is widely used to predict phenotypes in individuals. However, predictive models may fail to generalize to new datasets due to differences in population, data collection, and processing across datasets.
Goal(s): To resolve the dataset difference issue, we aimed to generalize knowledge from multiple diverse source datasets and translate the model to new target data.
Approach: Here we proposed Multi-domain and Uni-domain Fusion (MUF) method that combines cross-domain learning and intra-domain learning, to capture both domain-general information and domain-specific information.
Results: The results show that our MUF outperformed 4 strong baseline methods on 6 target datasets.
Impact: Our MUF method is adept at addressing the challenges introduced by different population profiles, fMRI processing pipelines, and prediction tasks. We offer a robust and universal learning strategy for domain-generalization in fMRI-based phenotypic prediction.
Introduction
Resting-state functional connectivity (RSFC) is widely used to predict phenotypes in individuals1,2. Due to unavoidable small sample size issues in neuroimaging studies3-7, there have been emerging works translating models trained from large-sized neuroimaging datasets to predicting phenotypes on small target datasets7,8. However, predictive models may fail to generalize to new datasets due to differences in population profiles, data collection, and processing pipelines across datasets9,10. Here we proposed a method named Multi-domain and Uni-domain Fusion (MUF) to enhance model generalizability across sites, which outperformed four strong baseline methods on six target datasets.Methods
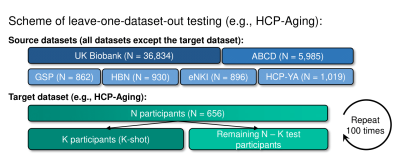
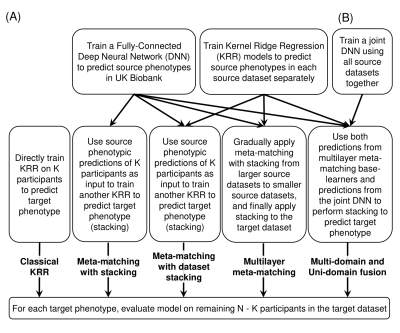
We used 419 by 419 Resting-state functional connectivity (RSFC) matrices to predict phenotypes in individuals7. Our study used 7 datasets: UK Biobank12,13, ABCD14, GSP15, HBN16, eNKI17, HCP-YA18 and HCP-Aging19. We did a leave-one-dataset-out test on each dataset (except UK Biobank since we need this only super-sized dataset for training): each time one dataset was used as the target dataset and all others were used as source datasets (Fig 1). Predictive models trained from source datasets were adapted to K participants (K-shot) in the target dataset to predict target phenotypes. The adapted models were evaluated in the remaining test participants. This procedure was repeated 100 times for stability. We compared MUF against 4 baseline methods (Fig 2A): classical Kernel Ridge Regression (KRR), meta-matching with stacking7, and our previous work: meta-matching with dataset stacking11 and multilayer meta-matching11.Classical KRR models were directly trained on K-shot to predict target phenotypes7. For meta-matching with stacking7, a feedforward deep neural network (DNN) was trained on the UK Biobank to predict 67 source phenotypes. The base DNN was applied to K-shot and DNN predictions were used as features to train a KRR model on K-shot to predict target phenotypes (i.e. stacking). Meta-matching with dataset stacking11 extends meta-matching with stacking7 by training separate KRR/DNN models for each source dataset and then performing stacking on K-shot to predict target phenotypes. To resolve the sample size imbalance across source datasets, multilayer meta-matching11 gradually applied stacking from larger source datasets to smaller source datasets to boost prediction accuracies. These predictive models then underwent another round of stacking using K-shot to predict target phenotypes.
Fig 2B shows our new proposed Multi-domain and Uni-domain Fusion (MUF) method. Notice that all the above 4 methods trained base models independently on each dataset. Thus, on the basis of multilayer meta-matching, we also employed a multi-domain learning strategy that trains a joint multi-task DNN using all source datasets together, enabling it to offset site differences to some extent and thus generalize better on unseen target datasets. Predictions from the above joint DNN were concatenated together with predictions from the multilayer meta-matching method and then used as features of the KRR stacking model to predict target phenotypes.
Results
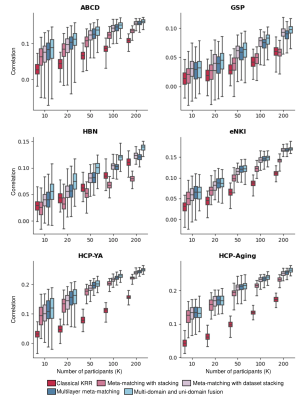
Fig 3 shows the prediction accuracy (Pearson's correlation) in six target datasets. All reported p values survived a false discovery rate of q < 0.05. Similar results were obtained with the coefficient of determination (COD).We found that original meta-matching with stacking7 was not even better than classical KRR on some datasets (e.g., HBN), but our proposed methods improved it by incorporating more diverse source datasets. In all test datasets, our MUF consistently outperformed all other baselines, indicating that MUF can better generalize from different source datasets and is more robust on new target datasets. After Wilcoxon signed-rank test, MUF was better than meta-matching with dataset stacking11 (p < 0.02 for all K values) and multilayer meta-matching11 (p < 0.02 for K > 20).
Discussion
Our MUF method is adept at addressing the challenges introduced by different population profiles, fMRI processing pipelines, and prediction tasks. Results show that our MUF enhanced the model generalization ability and boosted prediction performance on test datasets, by combining cross-domain learning and intra-domain learning to capture both domain-general information and domain-specific information. Overall, our MUF approach offers a robust and universal learning strategy for domain-generalization in fMRI-based phenotypic prediction.Conclusion
We propose a method named Multi-domain and Uni-domain Fusion (MUF), which uses both multi-domain and uni-domain learning strategies to translate phenotypic prediction models from multiple source datasets to small-sized target datasets. We found that our MUF performed the best on six test datasets.Acknowledgements
This work was supported by the Singapore National Research Foundation(NRF) Fellowship Class of 2017 (B.T.T.Y.), the NUS Yong Loo Lin School of Medicine NUHSRO/2020/124/TMR/LOA (B.T.T.Y.), the Singapore National Medical Research Council (NMRC) LCG OFLCG19May-0035 (B.T.T.Y.), the NMRC STaR20nov-0003 (B.T.T.Y.), the Healthy Brains Healthy Lives initiative from the Canada First Research Excellence Fund (D.B.), the Canada Institute for Advanced Research CIFAR Artificial Intelligence Chairs program (D.B.), Google Research Award (D.B.) and National Institutes of Health (NIH) R01AG068563A (D.B.), NIH R01MH120080 (A.J.H.) and NIH R01MH123245 (A.J.H.). Our computational work was partially performed on resources of the National Supercomputing Centre, Singapore (https://www.nscc.sg). Any opinions, findings and conclusions or recommendations expressed in this material are those of the author(s) and do not reflect the views of the Singapore NRF or NMRC.References
1. Eickhoff, S. B., & Langner, R. (2019). Neuroimaging-based prediction of mental traits: Road to utopia or Orwell?. PLoS biology, 17(11), e3000497.
2. Varoquaux, G., & Poldrack, R. A. (2019). Predictive models avoid excessive reductionism in cognitive neuroimaging. Current opinion in neurobiology, 55, 1-6.
3. Arbabshirani, M. R., Plis, S., Sui, J., & Calhoun, V. D. (2017). Single subject prediction of brain disorders in neuroimaging: Promises and pitfalls. Neuroimage, 145, 137-165.
4. Bzdok, D., & Meyer-Lindenberg, A. (2018). Machine learning for precision psychiatry: opportunities and challenges. Biological Psychiatry: Cognitive Neuroscience and Neuroimaging, 3(3), 223-230.
5. Masouleh, S. K., Eickhoff, S. B., Hoffstaedter, F., Genon, S., & Alzheimer's Disease Neuroimaging Initiative. (2019). Empirical examination of the replicability of associations between brain structure and psychological variables. elife, 8, e43464.
6. Poldrack, R. A., Huckins, G., & Varoquaux, G. (2020). Establishment of best practices for evidence for prediction: a review. JAMA psychiatry, 77(5), 534-540.
7. He, T., An, L., Chen, P., Chen, J., Feng, J., Bzdok, D., ... & Yeo, B. T. (2022). Meta-matching as a simple framework to translate phenotypic predictive models from big to small data. Nature Neuroscience, 1-10.
8. Lu, B., Li, H. X., Chang, Z. K., Li, L., Chen, N. X., Zhu, Z. C., ... & Yan, C. G. (2022). A practical Alzheimer’s disease classifier via brain imaging-based deep learning on 85,721 samples. Journal of Big Data, 9(1), 1-22.
9. Kakarmath, S., Esteva, A., Arnaout, R., Harvey, H., Kumar, S., Muse, E., ... & Kvedar, J. (2020). Best practices for authors of healthcare-related artificial intelligence manuscripts. NPJ digital medicine, 3(1), 134.
10. Abraham, A., Milham, M. P., Di Martino, A., Craddock, R. C., Samaras, D., Thirion, B., & Varoquaux, G. (2017). Deriving reproducible biomarkers from multi-site resting-state data: An Autism-based example. NeuroImage, 147, 736-745.
11. Chen, P., An, L., Wulan, N., Zhang, S., Ooi, Q. R., Kong, R., ... & Yeo, B. T. (2023). Multilayer meta-matching: translating phenotypic predictive models from multiple datasets with disparate sample sizes to small data. In Prep.
12. Sudlow, C., Gallacher, J., Allen, N., Beral, V., Burton, P., Danesh, J., ... & Collins, R. (2015). UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS medicine, 12(3), e1001779.
13. Miller, K. L., Alfaro-Almagro, F., Bangerter, N. K., Thomas, D. L., Yacoub, E., Xu, J., ... & Smith, S. M. (2016). Multimodal population brain imaging in the UK Biobank prospective epidemiological study. Nature neuroscience, 19(11), 1523-1536.
14. Volkow, N. D., Koob, G. F., Croyle, R. T., Bianchi, D. W., Gordon, J. A., Koroshetz, W. J., ... & Weiss, S. R. (2018). The conception of the ABCD study: From substance use to a broad NIH collaboration. Developmental cognitive neuroscience, 32, 4-7.
15. Holmes, A. J., Hollinshead, M. O., O’keefe, T. M., Petrov, V. I., Fariello, G. R., Wald, L. L., ... & Buckner, R. L. (2015). Brain Genomics Superstruct Project initial data release with structural, functional, and behavioral measures. Scientific data, 2(1), 1-16.
16. Alexander, L. M., Escalera, J., Ai, L., Andreotti, C., Febre, K., Mangone, A., ... & Milham, M. P. (2017). An open resource for transdiagnostic research in pediatric mental health and learning disorders. Scientific data, 4(1), 1-26.
17. Nooner, K. B., Colcombe, S. J., Tobe, R. H., Mennes, M., Benedict, M. M., Moreno, A. L., ... & Milham, M. P. (2012). The NKI-Rockland sample: a model for accelerating the pace of discovery science in psychiatry. Frontiers in neuroscience, 6, 152.
18. Van Essen, D. C., Smith, S. M., Barch, D. M., Behrens, T. E., Yacoub, E., Ugurbil, K., & Wu-Minn HCP Consortium. (2013). The WU-Minn human connectome project: an overview. Neuroimage, 80, 62-79.
19. Harms, M. P., Somerville, L. H., Ances, B. M., Andersson, J., Barch, D. M., Bastiani, M., ... & Yacoub, E. (2018). Extending the Human Connectome Project across ages: Imaging protocols for the Lifespan Development and Aging projects. Neuroimage, 183, 972-984.Figures