2217
Combining domain knowledge and foundation models for one-shot spine labeling1GE Healthcare, Bangalore, India, 2GE Healthcare, Niskayuna, NY, United States
Synopsis
Keywords: Analysis/Processing, Spinal Cord, MRI, Spine, Spine Labelling, Foundation Model, ML/AI
Motivation: Spine labelling is a step crucial for several important tasks such as MRI scan planning or associating image regions with mentions in clinical reports and others. Automating it can lead to significant benefits but developing automated solutions requires extensive annotations of vertebra labels.
Goal(s): To automate spine labelling without extensively training a DL model with manual annotations.
Approach: We adapted a vision foundational model-based approach that combines spine domain knowledge to predict spine labels.
Results: Our spine labelling method gives an average accuracy of 79% and 86% for cervical and lumbar high resolution T1 images, respectively.
Impact: Leveraging spatially relevant landmarks (disc) and vision foundation deep learning model, spine labels are predicted using one-shot localization. The proposed method doesn’t require any prior data for model training.
Introduction
Existing methods for spine labeling are based on using existing annotations to train deep learning (DL) models using data in range of 10,000 cases1-2. In recent times, foundation DL models pretrained on natural images, have shown amazing capacity to extract meaningful pixel-level features. These models based on ViTs and Stable Diffusion have been used for few shots localization and segmentation tasks3. The general methodology for this family of methods is to use the patch level features from ViT-based networks or intermediate level features from Stable Diffusion models and perform interpolation to obtain pixel level features for images. Given a reference template image and the corresponding pixel locations of regions of interest, a similar region of interest can be obtained on a target image, by performing a similarity search on the template image pixel features with features of all pixels in target image to identify the most similar region in the target image. Such a method, though effective for most tasks, may face a challenge while performing spine labelling due to multiple vertebrae regions with similar appearance. We propose to mitigate these issues by leveraging the relative position of vertebra and disc while assessing pixel similarity. This method requires user to mark the vertebrae labels on only a single template image/volume and then use this marking to scale spine labeling across any number of new spine test data.Methods
In this work, we present and evaluate two methods for spine labelling for cervical and lumbar T1 scans:a) Vanilla Approach: In this method for each vertebra pixel in the template image, the most similar image pixel from the target image is identified and labeled as the corresponding vertebra label. This is the default method for labelling with foundation model features3.
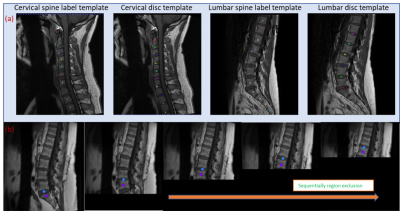
b) Sequential Conquer Approach (SCA): This novel method involves sequential labeling of vertebra by excluding image regions corresponding to vertebra already labeled. For this, a template vertebra labels, and disc region labels are generated (Figure 1a). The first vertebra identified is C1 for cervical and S1 for lumbar images using pixel similarity. Subsequently, the image region below the disc corresponding to S1 or above the disc corresponding to C1 is masked (pixel intensity set to 0), both in the template and the target image. The subsequent labeling is performed by searching for the most similar pixel with respect to template C1 in cervical and L5 in lumbar in the modified images, and the process is iterated till we reach C7 and L1 in cervical and lumbar, respectively. This process is shown in Figure 1b.
Evaluation: To evaluate the accuracy of spine detection we use a prediction scoring system. A spine label is considered correct (score of 1) if it falls within the vertebra within which the corresponding ground truth (GT) lies, else the score is 0. For all the labels in an image, an average score is then calculated, which is reported as accuracy of spine label prediction. Additionally, we report a distance metric, wherein we compute the Euclidean distance in mm between the predicted spine label coordinate and the corresponding GT spine label coordinate. We report the mean distance between the ground truth coordinates across all labels and all cases. We evaluated the two spine labelling approaches on a total of 90 T1W images, 57 cervical and 33 lumbar.
Results and Discussion
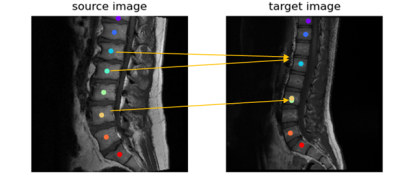
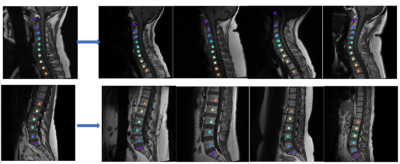
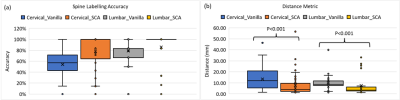
Figure 2 shows the results of the vanilla approach, wherein feature similarity of the spine vertebrae results in mislabeling. To address this issue, SCA was adopted. Using SCA, Figure 3 shows the sample labelling of the spine in several cases. It should be noted that if the first reference prediction (C1 or S1) goes wrong, mislabeling is inevitable. Furthermore, for any change in the target image such change of contrast, spine deformities, etc., the template image must be updated to match the changes. For both cervical and lumbar images (Figure 4a), our SCA method performed superior to vanilla (average accuracy of 79% versus 54% for cervical and 86% versus 79% for lumbar). This is further corroborated by significantly lower (p<0.001) distance metric for both the cases(Figure 4b).Conclusion
In this work, we present leveraging spatially relevant alternative landmarks such as disc surrounding the spine vertebrae labels to improve one-shot prediction of spine labels using foundational DL model. In case we need to support different data manifestation such as extreme scoliosis, we only need to change the template used for labelling and we could proceed to any specific imaging manifestation.Acknowledgements
No acknowledgement found.References
- Detection and Labeling of Vertebrae in MR Images Using Deep Learning with Clinical Annotations as Training Data - PubMed (nih.gov)
- 2-step deep learning model for landmarks localization in spine radiographs | Scientific Reports (nature.com)
- Deepa Anand et al., “One-shot Localization and Segmentation of Medical Images with Foundation Models”, 2023. https://doi.org/10.48550/arXiv.2310.18642
Figures