0726
Image Quality Assessment using an Orientation Recognition Network for Fetal MRI1Department of Biomedical Engineering, School of Medicine, Tsinghua University, Beijing, China, 2Tanwei College, Tsinghua University, Beijing, China
Synopsis
Keywords: Fetal, Brain, Data Analysis, Data Process, Image Reconstruction
Motivation: Fetal MRI is important in clinical and scientific applications but prone to motion artifacts. Automated image quality assessment (IQA) assists data acquisition and subsequent analyses. However, training neural networks for IQA requires labor-intensive manual annotation.
Goal(s): To develop a model for fetal MRI IQA that doesn't require image quality labels.
Approach: A network is trained to determine the acquisition orientation of 2D T2-weighted images. The variation of orientation recognition network (ORN) inferences for central images of a brain stack is used to assess motion and the image quality.
Results: High-quality and low-quality images are robustly discriminated. Image super-resolution from brain stacks is improved.
Impact: ORN-IQA eradicates the necessity image quality labels for training, thereby circumventing manual annotation. ORN-IQA simplifies online image quality evaluation and permits image reacquisition during fetal MR scans. Moreover, ORN-IQA improves super-resolution reconstruction results.
Introduction
MRI is crucial for assessing fetal brain development and pathology, owing to its superior tissue contrast compared to ultrasound imaging1–3. Nevertheless, the acquisition of 2D thick-slice T2-weighted images is vulnerable to inter- and intra-slice motion4 (Figure 1), resulting in reduced image quality and accuracy of subsequent analyses (e.g., super-resolving 3D isotropic resolution image volumes from axial, sagittal and coronal image stacks for brain morphometry).Recent efforts employ deep learning models for image quality assessment (IQA) in fetal MRI4–8, aiming for on-the-fly IQA and image re-acquisition during scans. However, most methods require image quality labels, obtained through time-consuming and subjective visual inspection by expert radiologists6. The cost of training such models is prohibitively high.
To address this challenge, we propose an orientation recognition network (ORN) for quality-label-free IQA (ORN-IQA, Fig. 2). Essentially, ORN-IQA trains a neural network to discriminate axial, coronal, and sagittal orientation of central images of a brain image stack and quantifies the uncertainty of ORN predicts for IQA. In the case of large motion, image quality decreases, leading to inconsistent predictions and thus elevated variation. In contrast to quality labels, orientation labels can be freely obtained from the image file header information9. The efficacy of ORN-IQA for classifying high and low-quality images and improving isotropic image reconstruction and segmentation is demonstrated.
Methods
Data Acquisition and Pre-processing (Fig. 2A). Seventy-four pregnant women (20-36 weeks gestation) with normal fetal brains were enrolled with informed written consent and IRB approval to obtain 2D T2-weighted images, along axial, coronal, and sagittal directions using a turbo spin echo (TSE) sequence. For low-quality stacks, repeated data might be acquired determined by the technician. The super-resolution reconstruction method NiftyMIC10 was employed to perform slice-to-volume motion correction and reconstruct a single fetal brain volume at 0.8 mm isotropic spatial resolution from stacks of 2D images acquired along axial, sagittal and coronal directions. Sixty cases were selected as the training set (288 stacks) and validation set (72 stacks), with the remaining 14 cases (83 stacks) reserved for testing. Brain masks were obtained from 2D TSE images using a public fetal brain localization network10.Network Architecture (Fig. 2B). A deep learning model called ORN was proposed for classifying TSE images into axial, coronal, and sagittal orientations. The network architecture followed the SFCN11. The first five blocks were employed to extract features from the input. The sixth block increased non-linearity of the model. The final block mapped features to classification vectors.
Training and Testing Pipeline (Fig. 2C). The central seven image slices within the range of [⌈N/2⌉-3, ⌈N/2⌉+3] were used as the network input for training and validation (N ranged between 12 to 36). Each image was randomly rotated for data augmentation during the training. The validation set was used to assess whether the network has converged. Training was halted when the validation accuracy stabilized.
During the testing, the central seven slices of each brain stack were fed into the ORN, yielding seven predictive vectors $$$\left\{p_{0}, p_{1}, \ldots, p_{6}\right\}$$$. The j-th predictive vector $$$p_{j}=\left\{p_{j, 1}, p_{j, 2}, p_{j, 3}\right\}$$$ were the probabilities of the (⌈N/2⌉-3+j)-th slice belonging to each orientation. Finally, the variation of the prediction results was calculated as the entropy:
$$ Q_{s}=-\sum_{i}\left(\frac{1}{n} \sum_{j} p_{j, i}\right) \log \left(\frac{1}{n} \sum_{j} p_{j, i}+\epsilon\right) $$
where $$$\epsilon$$$ is an extremely small positive value used to prevent logarithmic calculation errors.
Results
The IQA results show that high-quality images consistently yield very low $$$Q_s$$$ scores, while low-quality images yield high $$$Q_s$$$ scores (Fig. 3). In row eight (highlighted in red boxes), the slice changes from sagittal orientation to coronal orientation, leading to increased $$$Q_s$$$. A slice-wise IQA model8 can't detect such inter-slice motion as both individual slices exhibit high quality.For twenty-one cases with re-acquired image stacks, ORN-IQA retrospectively classified 85.7% initially acquired data into lower quality, which was consistent with the fact that additional repeats needed to be acquired (Fig. 4A).
ORN-IQA effectively prevented the negative influence of low-quality brain stacks for subsequent analyses by filtering them out. In cases 1 and 2, ORN-IQA excluded a coronal and axial stack respectively, facilitating NiftyMIC super-resolution reconstruction and brain segmentation12 (Fig. 5A). Without ORN-IQA, NiftyMIC reconstruction using low-quality stacks was wrong, inducing errors in subsequent brain segmentation and morphometry (Fig .5B).
Discussion
Fig. 4B demonstrates orientation recognition accuracy results for using different number of image slices, with seven or nine slices achieving the highest validation accuracy of 97.22%.Conclusion
ORN-IQA can be used for automatically determining whether repeats should be re-acquired and for selecting high-quality image stacks for NiftyMIC reconstruction. Future work will deploy ORN-IQA for clinical applications.Acknowledgements
Funding was provided by the Tsinghua University Startup Fund.References
1. Manganaro L, Capuani S, Gennarini M, et al. Fetal MRI: what’s new? A short review. Eur Radiol Exp. 2023;7(1):41. doi:10.1186/s41747-023-00358-5
2. Fileva N, Severino M, Tortora D, Ramaglia A, Paladini D, Rossi A. Second trimester fetal MRI of the brain: Through the ground glass. J of Clinical Ultrasound. 2023;51(2):283-299. doi:10.1002/jcu.23423
3. Huang Y, Liu G, Luo Y, Yang G. ADFA: Attention-Augmented Differentiable Top-K Feature Adaptation for Unsupervised Medical Anomaly Detection. In: 2023 IEEE International Conference on Image Processing (ICIP). IEEE; 2023:206-210. doi:10.1109/ICIP49359.2023.10222528
4. Lim A, Lo J, Wagner MW, Ertl-Wagner B, Sussman D. Automatic Artifact Detection Algorithm in Fetal MRI. Front Artif Intell. 2022;5:861791. doi:10.3389/frai.2022.861791
5. Liao L, Zhang X, Zhao F, et al. Joint Image Quality Assessment and Brain Extraction of Fetal MRI Using Deep Learning. In: Martel AL, Abolmaesumi P, Stoyanov D, et al., eds. Medical Image Computing and Computer Assisted Intervention – MICCAI 2020. Vol 12266. Lecture Notes in Computer Science. Springer International Publishing; 2020:415-424. doi:10.1007/978-3-030-59725-2_40
6. Largent A, Kapse K, Barnett SD, et al. Image Quality Assessment of Fetal Brain MRI Using Multi‐Instance Deep Learning Methods. Magnetic Resonance Imaging. 2021;54(3):818-829. doi:10.1002/jmri.27649
7. Sanchez T, Esteban O, Gomez Y, Eixarch E, Cuadra MB. FetMRQC: Automated Quality Control for Fetal Brain MRI. In: Link-Sourani D, Abaci Turk E, Macgowan C, Hutter J, Melbourne A, Licandro R, eds. Perinatal, Preterm and Paediatric Image Analysis. Vol 14246. Lecture Notes in Computer Science. Springer Nature Switzerland; 2023:3-16. doi:10.1007/978-3-031-45544-5_1
8. Xu J, Lala S, Gagoski B, et al. Semi-supervised Learning for Fetal Brain MRI Quality Assessment with ROI Consistency. In: Martel AL, Abolmaesumi P, Stoyanov D, et al., eds. Medical Image Computing and Computer Assisted Intervention – MICCAI 2020. Vol 12266. Lecture Notes in Computer Science. Springer International Publishing; 2020:386-395. doi:10.1007/978-3-030-59725-2_37
9. Ison M, Dittrich E, Donner R, Kasprian G, Prayer D, Langs G. Fully Automated Brain Extraction and Orientation in Raw Fetal MRI. Published online 2012. doi:10.13140/2.1.2312.8966
10. Ebner M, Wang G, Li W, et al. An automated framework for localization, segmentation and super-resolution reconstruction of fetal brain MRI. NeuroImage. 2020;206:116324. doi:10.1016/j.neuroimage.2019.116324
11. Peng H, Gong W, Beckmann CF, Vedaldi A, Smith SM. Accurate brain age prediction with lightweight deep neural networks. Medical Image Analysis. 2021;68:101871. doi:10.1016/j.media.2020.101871
12. Fidon, Lucas, et al. "A Dempster-Shafer approach to trustworthy AI with application to fetal brain MRI segmentation." arXiv preprint arXiv:2204.02779 (2022).
Figures
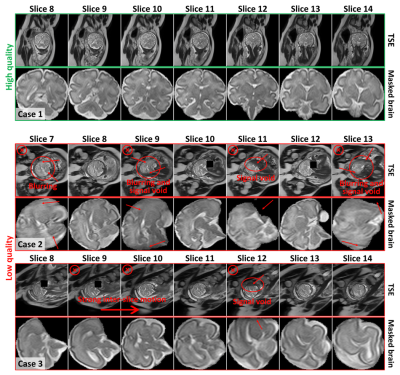
Figure 1. Examples of clinical image quality. Top, high quality 2D TSE T2-weighted images, and bottom, low quality TSE images. High quality images (Case 1) exhibit clear demarcation of brain structures, whereas low quality images (Case 2 and Case 3) display artifacts obscuring these features. Low quality image stacks would be deemed unsuitable for further analysis due to prominent intensity variations across numerous slices and pronounced signal attenuation, resulting from signal void and blurring over the brain region.
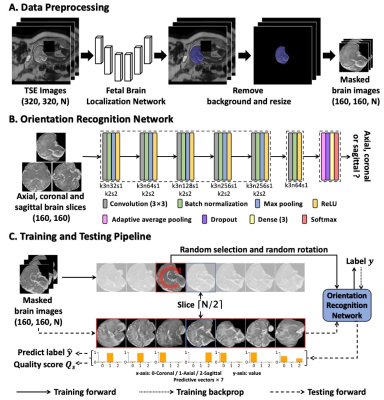
Figure 2. Proposed ORN-IQA method. (A) Data preprocessing involves brain extraction using a network, followed by background removal and resizing. (B) The ORN takes a brain slice as input and produces the predictive vector. The abbreviation k3n32s1 for convolution layer means kernel size of 3×3, the kernel number of 64 and the stride of 1. The k2s2 is for max pooling layer. (C) In the training phase, a randomly selected slice from the range of [⌈N/2⌉-3, ⌈N/2⌉+3] with random rotation for training. In the testing phase, seven slices are used as input to generate predictive vectors.
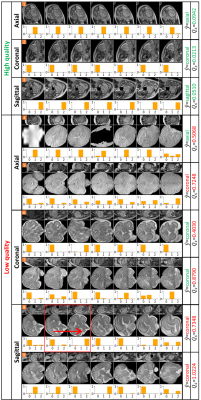
Figure 3. Image quality assessment results. The top three rows show IQA results for high quality TSE images. The following rows (fourth to ninth) display IQA results for low quality images. $$$\hat{y}$$$ represents ORN’s predicted orientation, with red/green indicating incorrect/correct predictions. $$$Q_s$$$ is the quality score derived from each predictive vector. Higher $$$Q_s$$$ signifies lower image quality, and $$$\hat{y}$$$ for low-quality images may be incorrect. Red/green indicates the $$$Q_s$$$ below/above the exclusion threshold which is set to 0.2.
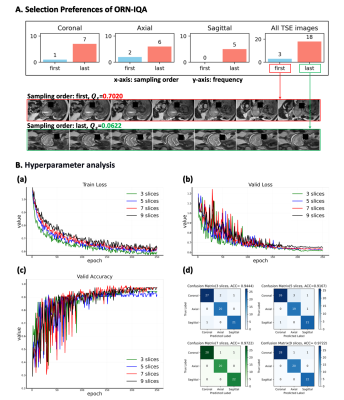
Figure 4. Further analysis. (A) The technician typically re-acquired images due to suboptimal initial MR quality. ORN-IQA retrospectively classified 85.7% initially acquired data into lower quality for twenty-one cases with re-acquired image stacks. (B) In ORN-IQA, the number of image slices is a hyperparameter. ORN converges post 250 epochs, indicated by stable training and validation losses, while validation accuracy remains consistent (a, b, c). With seven or nine slices, peak validation accuracy is 97.22%. To reduce IQA time-consuming, we selected seven slices (d).
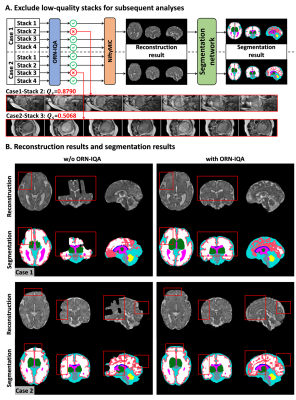
Figure 5. Improvement in reconstruction results. (A) Proposed ORN-IQA can exclude low-quality brain stacks to prevent them from affecting subsequent analyses. For case 1 and case 2, ORN-IQA excluded a coronal-oriented stack and an axial-oriented stack, respectively. The remaining high-quality stacks are used for NiftyMIC reconstruction and AI segmentation. (B) Without using ORN-IQA, NiftyMIC is affected by low-quality stacks, leading to reconstruction failure and further affecting segmentation and subsequent brain morphometry.