0723
Radio-pathomic maps of complex histo-morphometric features trained with whole mount prostate histology differentiate prostate cancer on MPMRI1Biophysics, Medical College of Wisconsin, Milwaukee, WI, United States, 2Medical College of Wisconsin, Milwaukee, WI, United States, 3University of California - Davis, Sacramento, CA, United States
Synopsis
Keywords: Prostate, Body
Motivation: The motivation of this study is to develop novel methods for mapping non-invasively the underlying history-morphometric features of prostate cancer.
Goal(s): The goal of this study is to develop and demonstrate radio-pathomic mapping techniques to enable noninvasive detection of prostate cancer presence and distinction from benign tissue using MRI.
Approach: Our approach was to align multiparametric MRI with digitized histology slides from prostatectomy specimens, then predict quantitative histological features from MRI intensities, and use these predicted features to classify cancer versus noncancer regions.
Results: Our models can distinguish cancerous from noncancerous prostate tissue with 70% accuracy.
Impact: This study uses radio-pathomic mapping for noninvasive prostate cancer detection, demonstrating the potential to differentiate cancerous vs benign prostate tissue using imaging surrogates of microstructural features discernible only on histology.
Introduction
Prostate cancer (PCa) is the most diagnosed male cancer in the United States accounting for 1 in 8 new cancer diagnoses. Multiparametric MRI (MP-MRI), including T2-weighted imaging (T2WI) and apparent diffusion coefficient (ADC) maps calculated from diffusion weighted imaging (DWI), has been used to detect PCa, and has been a promising tool in diagnosing high-grade PCa. The Prostate Imaging Reporting and Data System (PI-RADS) has standardized acquisition and reporting of prostate MRI. PI-RADS has aided in the detection of cancerous lesions and has improved consistency of clinical radiology reads. The underlying pathology however remains a mystery until biopsy or radical prostatectomy. Radio-pathomic maps have been advantageous in detecting cancer on MRI, including in glioblastoma patients[1-3]. Therefore, this study tested the hypothesis that radio-pathomic maps of prostate cancer could predict cancer presence on conventional MR imaging.Methods
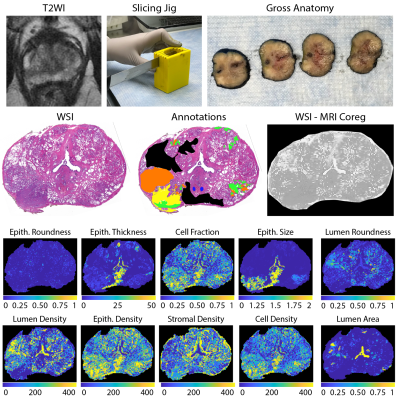
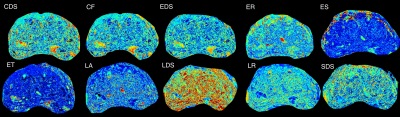
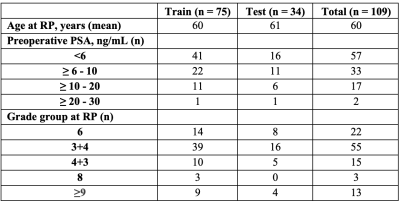
A total of 109 patients with pathologically-confirmed prostate cancer were recruited for this study. Demographic features are presented in Table 1, and patients were split into a 2/3 – 1/3 train/test cohort, balanced by Gleason Grade Group. Patients underwent MP-MRI prior to surgery on a 3T MRI scanner with an endorectal coil. Each protocol included T2W, DWI, and resulting ADC maps. All T2W were intensity normalized using the z-score of intensity within the mask of the prostate. Robotic prostatectomy was performed following imaging approximately 4 weeks after imaging. Prostate samples were formalin fixed overnight and sectioned using a custom, 3D printed slicing jig modeled after the orientation and slice thickness of the T2W. Briefly, prostate masks were manually drawn and 3D modeled using to create patient-specific tissue slicing jigs (Figure 1). Whole-mount tissue sections were paraffin-embedded, axially sectioned, hematoxylin and eosin (H&E) stained, and digitized at 40x magnification. Scanned slides were annotated by a GU pathologist to identify regions of unique Gleason patterns, including Gleason 3 (G3), G4 non-cribriform glands (G4NC), G4 cribriform (to papillary) glands (G4CG), and Gleason 5 (G5), as well as non-cancerous regions including seminal vesicles, atrophy, and high-grade prostatic intraepithelial neoplasia (HGPIN). All cancerous annotations were combined into one class for these analyses. An example of these annotations can be found in Figure 1. A color deconvolution algorithm was applied to the H&E stained slides, and the resulting intensity standardized images were then computationally processed to segment tissue into lumen, stroma, epithelium, and epithelial cell masks. These masks were used to calculate first-order features (FOF) (i.e., their respective densities), as well as second-order histo-morphometric features (SOF) including lumen roundness and area; epithelium roundness, area, thickness, size; and cell fraction (i.e., percent nuclei per gland epithelium). Slide, annotations, and histo-morphometric features were nonlinearly registered to the T2WI using in house control-point based software (n = 3 slides/patient). Histo-morphometric features as well as T2WI and ADC intensities were averaged across annotation per lesion and unlabeled tissue. Radio-pathomic models were then trained using the histological and MR intensity features. First, a bagged ensemble regression was fitted for each individual feature using T2WI and ADC intensities as input to non-invasively predict these histological features in MR space. Next, the histological features individually, as well as combinations of FOF, SOF, and all histological features were used as input to a bagged ensemble machine learning model to predict cancer presence. Models were first trained and tested with our ground-truth features. Final testing was completed on the cancer presence models using the predicted output from each regression.Results
Regression models to predict histological features using MRI intensity values were tested using RMSE. Models were considered to have good performance if the RMSE fell within an standard deviation of the test set feature. All regressions successfully were able to predict their features. From our cancer model, cell density (CDS), FOF, and SOF were all able to predict cancer from noncancer with an overall accuracy of 70%, with each model additionally predicting both classes at 70% accuracy.Discussion
These results suggest that radio-pathomic mapping of prostate cancer may provide non-invasive insight into cancer presence. MRI intensity values were able to accurately predict histo-morphometric features at RMSEs within a standard deviation of the feature, and predictions from the regressions were able to classify cancer presence with an overall accuracy of 70%. Future studies are warranted to determine if these features can additionally predict high from low-grade cancer and unique Gleason patterns.Conclusion
This study demonstrated in a cohort of 109 prostate cancer patients that radio-pathomic mapping could accurately predict cancer presence, using MR intensity as input to feature models, and features to finally predict cancer.Acknowledgements
R01CA218144, R01CA249882References
1. Bobholz SA, Lowman AK, Brehler M, et al. Radio-Pathomic Maps of Cell Density Identify Brain Tumor Invasion beyond Traditional MRI-Defined Margins. AJNR Am J Neuroradiol. May 2022;43(5):682-688. doi:10.3174/ajnr.A74772.
2. McGarry SD, Bukowy JD, Iczkowski KA, et al. Radio-pathomic mapping model generated using annotations from five pathologists reliably distinguishes high-grade prostate cancer. J Med Imaging (Bellingham). Sep 2020;7(5):054501. doi:10.1117/1.JMI.7.5.0545013.
3. McGarry SD, Hurrell SL, Iczkowski KA, et al. Radio-pathomic Maps of Epithelium and Lumen Density Predict the Location of High-Grade Prostate Cancer. Int J Radiat Oncol Biol Phys. Aug 1 2018;101(5):1179-1187. doi:10.1016/j.ijrobp.2018.04.044
Figures