0722
Deep learning-based super-resolution imaging for routine clinical T1- and T2-weighted breast MRI at 1.5T1Philips GmbH Market DACH, Hamburg, Germany, 2Philips, Best, Netherlands, 3Diagnostic and Interventional Radiology, University Hospital RWTH Aachen, Aachen, Germany, 4Philips Japan, Tokyo, Japan
Synopsis
Keywords: Breast, Cancer, deep learning; super resolution; screening
Motivation: High-resolution images in breast MRI are desired for lesion detection and characterization but are restricted due to scan time constraint in routine clinical settings.
Goal(s): Our goal was to use deep learning (DL)-based reconstructions to improve image resolution and quality of routine clinical breast MRI.
Approach: We applied a dedicated Precise-Image-Net for both 2D T1- and T2-weighted imaging in breast cancer patients at 1.5T and compared it to conventional parallel imaging, compress sensing, and convolutional neural network (CNN) reconstructions.
Results: Initial clinical data demonstrated a clear improvement of sharpness in breast T1- and T2-weighted images compared with standard reconstructions.
Impact: Deep learning-based super-resolution reconstruction provides improved image resolution and sharpness in breast MRI, showing promises for better lesion detection and characterization in routine clinical settings without prolonging scan time, which is of particular importance in dynamic contrast enhanced-MRI.
Introduction
Breast MRI plays a vital role in lesion detection and characterization, particularly for breast cancer imaging. T1- and T2-weighted imaging are most often performed and serves as the fundamental part in breast MRI1,2. Though a high spatial resolution is desired to achieve its clinical purpose, it is inherently limited by examination time in routine clinical settings, even with the use of various scan acceleration techniques. In very recent years there has been a growing interest in applying Artificial Intelligence (AI), in particular, deep learning convolutional neural networks (CNN) to improve spatial resolution of medical images3-5. In this study, we investigated the use of prototype deep learning-based reconstructions to improve the image resolution of clinical T1- and T2-weighted breast images. Image quality was compared between conventional parallel imaging, compressed sensing, and two types of deep learning-based reconstructions.Methods
A total of 8 patients referred for breast MRI were examined on a 1.5T whole-body clinical system (Ingenia Ambition X, Philips Healthcare) using a 16-channel Sentinelle breast coil. Bilateral axial multi-slice 2D T2-TSE and T1-FFE sequences were acquired. Key scan parameters are listed in Table 1. For image reconstruction, the following techniques were applied to T2 before Gadolinium injection (pre-Gd) as well as pre- and post-Gd T1-weighted images: 1) conventional parallel imaging with sensitivity encoding (SENSE)6; 2) regularized, iterative L1-norm minimization after wavelet sparsifying transformation by combining compressed sensing and SENSE coil information (C-SENSE)7-9; 3) Adaptive-CS-Net for noise and artifacts reduction based on the same C-SENSE undersampled data (CSAI)10,11; 4) Precise-Image-Net, also known as SuperResolution network, for further removal of the noise and ringing artifacts (SupRes)4,5,12. Specifically, Adaptive-CS-Net allows to reconstruct images acquired with Compressed-SENSE based on variable density undersampling patterns. This CNN is applied during coil combination, to remove noise and undersampling-related artifacts. Precise-Image-Net is applied to replace the traditional zero-filling approach to increase the reconstructed matrix size and therewith to improve the sharpness of the images. This network has been trained on pairs of low- and high-resolution data with k-space crops to induce ringing artifacts. In addition, data consistency checks were implemented to match the resulting k-space with the measured k-space data. The full reconstruction pipeline generates images with a higher matrix size and reduced ringing artefacts for 2D Cartesian acquisitions12.Two AI/DL-based image reconstruction techniques consisting of a series of convolutional neural networks (CNN) were available in the implemented prototype software (Philips SmartSpeed and PreciseImage). All images were reconstructed on the console computer.
Image quality was compared between all four reconstruction techniques by means of visual inspection and by means of signal profile and histogram analysis using an in-house developed Python program.
Results and Discussion
Data were acquired from 8 female patients (average age 51.9 ± 14.3 years). In general, overall image quality was found much improved in images reconstructed with the SupRes AI/DL technique, with higher signal strength and clearly higher sharpness of the tissue structures for both multi-slice 2D T2-TSE and T1-FFE images. Figure 1 shows T2-weighted images comparing SENSE, C-SENSE, CSAI, and SupRes. Visually, SupRes Precise-Image-Net images showed improved sharpness. Figure 2 shows T1-weighted images comparing SENSE, C-SENSE, CSAI, and SupRes, before and after contrast injection. In both T1 and T2-weighted contrasts, a higher reconstruction in-plane spatial resolution allowed SupRes images to better delineate the mammary anatomy. This resulted in sharper tissue structures compared to the other reconstructions, which used zero-filling to interpolate the image resolution. C-SENSE, CSAI, and SupRes (same acquired dataset) demonstrated comparable signal profiles. Noteworthy, in most cases high signal (e.g., from lipid) in the middle of the low-signal (e.g., glandular tissue) structures can be better depicted in SupRes. One case is shown in Figure 3, indicating improved image sharpness due to higher in-plane spatial resolution by Precise-Image-Net (aka SuperResolution network). Histogram analysis demonstrated comparable results between different reconstructions, indicating similar distribution of the pixel i.e., signal intensity values (Figures 4). Quantitative comparison was not reported due to low case number and statistical power. A detailed study to compare the effect and clinical performance of different reconstruction techniques is warranted.Conclusion
Deep learning-based super-solution reconstruction can improve image resolution and quality in routinely acquired breast MRI. Initial clinical data demonstrated clearly improved sharpness of both T1 and T2-weighted images before and after Gd injection, compared to the conventional reconstruction techniques. Further investigations are needed to validate its performance in larger cohorts.Acknowledgements
The authors thank Ms. Katja Berkmann and Ms. Ilse Rubie (Philips), Ms. Lucia Noël and Ms. Anna Fischer (Univ. Hospital RWTH Aachen) for assistance in preparation and patient data collection.References
1. Kuhl CK, Schrading S, Strobel K, et al. Abbreviated Breast Magnetic Resonance Imaging (MRI): First Postcontrast Subtracted Images and Maximum-Intensity Projection – A Novel Approach to Breast Cancer Screening With MRI. J Clin Oncol 32:2304. DOI: 10.1200/JCO.2013.52.5386.
2. Bode M, Huck LC, Zhang S, et al. Clinical evaluation of cylindrical regional suppression in dynamic contrast-enhanced breast MRI: An intra-individual comparison study on image quality and lesion conspicuity. Eur J Radiol 2023;161:110724. DOI: 10.1016/j.ejrad.2023.110724.
3. Li Y, Sixou B, Peyrin F. A Review of the Deep Learning Methods for Medical Images Super Resolution Problems. Irbm 2021;42(2):120-133. doi:10.1016/j.irbm.2020.08.004.
4. Kwon J, Yuda K, Yoneyama M, et al. Deep learning-based image quality and spatial resolution improvement for diffusion-weighted imaging in liver. ISMRM 2023, p3443, Toronto.
5. Kravchenko D, Isaak A, Mesropyan N, et al. AI-assisted upscaling of low-resolution cardiac MRI reduces acquisition time and yields qualitatively comparable images to standard techniques. ISMRM 2023, Toronto.
6. Pruessmann KP, Weiger M, Scheidegger MB, et al. SENSE: Sensitivity Encoding for Fast MRI. Magn Reson Med 1999;42:952.
7. Lustig M, Donoho D, and Pauly JM. Sparse MRI: The Application of Compressed Sensing for Rapid MR Imaging. Magn Reson Med 2007;58:1182.
8. Meister RL, et al. Compressed SENSE in Pediatric Brain TumorMR Imaging: Assessment of Image Quality, Examination Time and Energy Release. Clin Neuroradiol 2022. DOI: 10.1007/s00062-021-01112-3.
9. Bode M, Zhang S, Terwolbeck MN, et al. Liver diffusion‑weighted MR imaging with L1‑regularized iterative sensitivity encoding reconstruction based on single‑shot echo‑planar imaging: initial clinical experience. Scientific Reports 2022;12:12468. DOI: 10.1038/s41598-022-16324-x.
10. Pezzotti N, Yousefi S, Elmahdy MS et al. An adaptive intelligence algorithm for undersampled knee MRI reconstruction. IEEE Access 2020;8:204825.
11. Feuerriegel CG, Weiss K, Kronthaler S, et al. Evaluation of a deep learning‑based reconstruction method for denoising and image enhancement of shoulder MRI in patients with shoulder pain. European Radiology 2023. DOI: 10.1007/s00330-023-09472-9.
12. Bischoff LM, Peeters JM, Weinhold L, et al. Deep Learning Super-Resolution Reconstruction for Fast and Motion-Robust T2-weighted Prostate MRI. Radiology 2023, 308(3):e230427. DOI: 10.1148/radiol.230427.
Figures
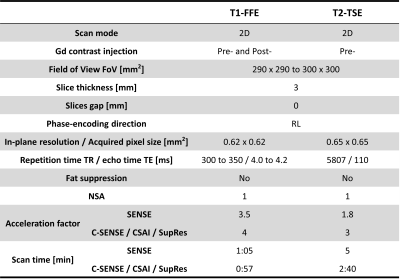
Table 1. Imaging parameters of the breast MRI pulse sequences used in this study.
FFE = fast field echo; TSE = turbo-spin echo; NSA = number of averages; SENSE = sensitivity encoding; C-SENSE = compressed SENSE; CSAI = Compressed SENSE AI; SupRes = Super Resolution.
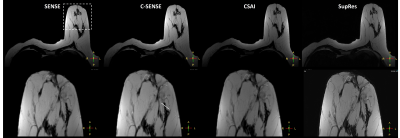
Figure 1. 2D T2-TSE images in a breast patient. The comparison shows different image reconstruction using conventional parallel imaging SENSE, C-SENSE, CSAI, and SupRes. CSAI and SupRes are DL-based CNNs (from left to right). Dotted box indicates zoomed view (bottom). The white line provides a signal profile plotted in Figure 3.
Details see text. TSE = turbo-spin echo; SENSE = sensitivity encoding; C-SENSE = compressed SENSE; CSAI = Compressed SENSE AI; SupRes = Super Resolution; DL = deep learning.
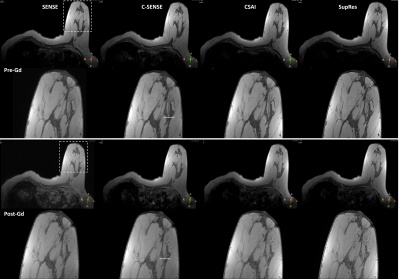
Figure 2. 2D T1-FFE images in a breast patient before (pre-Gd) and after (post-Gd) contrast medium administration. Comparison shows different image reconstruction using conventional parallel imaging SENSE, C-SENSE, CSAI, and SupRes. CSAI and SupRes are DL-based CNNs (from left to right). The dotted box indicates zoomed view (bottom). The white line provides a signal profile plotted in Figure 3. Details see text.
FFE = fast field echo; SENSE = sensitivity encoding; C-SENSE = compressed SENSE; CSAI = Compressed SENSE AI; SupRes = Super Resolution; DL = deep learning.
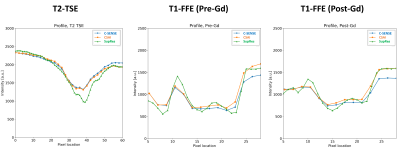
Figure 3. Comparison of signal profiles between C-SENSE, CSAI, and SupRes reconstructions in breast MRI. CSAI and SupRes are DL-based. Signal profile curves are plotted based on the white lines shown in Figure 1 and 2 for 2D T1-FFE and T2-TSE. Details see text.
FFE = fast field echo; TSE = turbo-spin echo; C-SENSE = compressed SENSE; CSAI = Compressed SENSE AI; SupRes = Super Resolution; DL=deep learning.
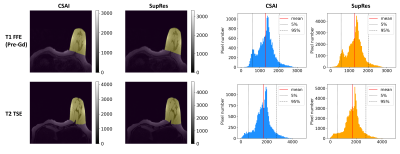
Figure 4. Histogram showing the pixel values comparing CSAI and SupRes DL-based reconstructions. Manual segmentations were obtained for the left breast (left), from which pixel values were extracted to create the histograms (right). Average values (red lines) as well as 5 and 95 percentiles (dotted lines) were indicated. Details see text.
FFE = fast field echo; TSE = turbo-spin echo; CSAI = Compressed SENSE AI; SupRes = Super Resolution; DL = deep learning.