0718
Deep learning model based on multiparametric MRI for prediction of synchronous liver metastasis from rectal cancer: a two-center study.1Clinical Oncology School of Fujian Medical University, Fuzhou, China, 2GE Healthcare, MR Research China, Beijing, China
Synopsis
Keywords: Cancer, Cancer, Rectal cancer, Deep learning radiomics, Magnetic resonance imaging, Synchronous liver metastasis
Motivation: Accurate synchronous liver metastasis (SLM) risk stratification is important for treatment planning and prognosis improvement.
Goal(s): Our goal is to establish a non-invasive and quantitative prediction model of synchronous liver metastases (SLM) in rectal cancer (RC) to help with accurate staging.
Approach: The deep learning (DL) model was fitted based on multi-parameter MRI of primary cancer combined with Clinical features (CF) features, and 5-fold cross-validation and external validation were performed.
Results: We demonstrated that the combination of CF and DL features achieved a satisfactory predictive performance for SLM, and also confirmed the generalizability of this model by external validation.
Impact: The discovery of the DL model would change treatment strategies. For patients with high-risk metastasis, a more aggressive systemic examination and shorter follow-up should be considered and may contribute to improved outcomes.
Background
Rectal cancer (RC) is one of the most common cancers of rectum and the leading cause of cancer-related deaths [1, 2]. The 5-year survival rate is only 14% in rectal cancer presenting with metastatic disease [3]. Approximately 20–34% of patients are diagnosed with synchronous liver metastasis (SLM), and 50–60% of patients will develop LM throughout their course [4]. Direct surgery for isolated metastatic lesions can achieve a more satisfactory 5-year survival rate [5], but only 25% of patients are suitable for surgery at diagnosis [6]. Therefore, early screening for SLM from rectal cancer in the high-risk group at preoperative examination is important to improve prognosis.Medical imaging can provide information regarding the entire tumor volume noninvasively. Commonly used methods for the detection and localization of SLM include CT, MRI, and PET-CT. These imaging modalities rely primarily on qualitative features. Recent advances in development of deep learning (DL) allows conversion of qualitative images into meaningful data. It can automatically learn crucial information from raw image data to perform tasks such as detection, classification, and segmentation [7]. Early results have demonstrated DL have good accuracy in the prediction of response to chemotherapy, distant metastasis, and prognosis in rectal cancer [8] [9].
Therefore, here we aimed to establish and validate DL models based on pre-treatment primary cancer multiparametric MRI images in noninvasively prediction of SLM in patients with rectal cancer.
Methods
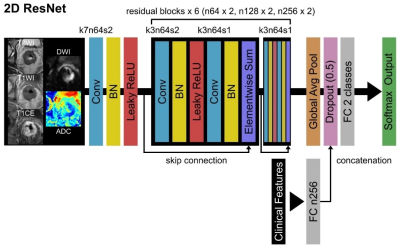
Institutional review board approval was obtained from Fujian Cancer Hospital and Fujian Medical University Union Hospital. 152 and 31 patients were included in primary and external validation cohorts, respectively. Clinical characteristics, including age, gender, and location of the lesion, CEA and CA199 levels were recorded. MRI examinations were performed on a 3.0T MRI scanner (Signa HDxt; GE Healthcare, Milwaukee, WI). Axial T1WI, T2WI, DWI, and T1CE were acquired. ADC maps were further calculated from DWI images.All data from internal study cohort were randomly divided into training and validation datasets in an 8:2 ratio. For image preprocessing, we performed rigid registration, resampling, and data augmentation, using MATLAB 2018a (MathWorks, Natick, MA). DL model was constructed based on the residual network published previously [10] (Figure 1). Two DL models were constructed with or without CF incorporation. Finally, the SLM prediction model was validated by 5-fold cross-validation and external validation. Statistical analysis of clinical characteristics was performed using SPSS 22.0 (IBM Corp, Armonk, NY). For the comparison of patient characteristics, categorical and continuous data were compared using Chi-square and Student t test, respectively. The performance of the model in predicting SLM was evaluated by ROC analysis, and AUC, ACC, SEN, SPE, PPV, and NPV were recorded.
Results
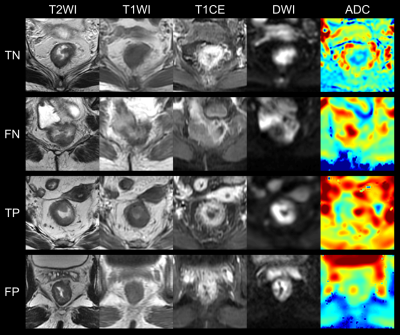
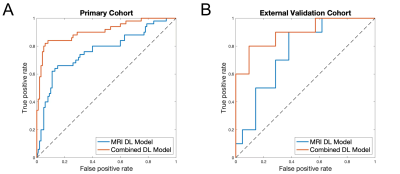
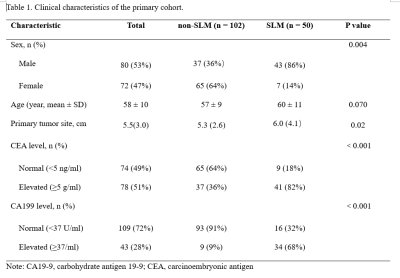
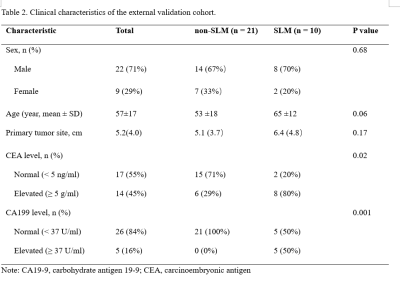
50 of 152 cases and 10 of 31 cases had SLM diagnosed by MRI or CT in primary and external validation cohorts, respectively. Detailed clinicopathological characteristics and comparison results were summarized in Table 1 and 2. In the primary cohort, there was statistic difference in sex, primary tumor site, CEA, and CA199 between two groups. In the external validation cohort, CEA and CA199 were significantly different between two groups.Figure 2 shows representative input images and the successful and unsuccessful predictions by the DL model. Figure 3A shows ROC curves in discriminating two groups in the primary study cohort. The DL model without CF and the combined model achieved AUCs of 0.765 (95% CI: 0.679, 0.851) and 0.909 (95% CI: 0.853, 0.964), respectively. For the combined DL model, the ACC, SEN, SPE, PPV, and NPV were 0.893, 0.840, 0.920, 0.840, and 0.920, respectively. Figure 3B shows ROC curves in discriminating two groups in the external validation cohort. The DL model without CF and the combined model achieved AUCs of 0.757 (95% CI: 0.587, 0.928) and 0.895 (95% CI: 0.770, 1.000), respectively. For the combined model, the ACC, SEN, SPE, PPV, and NPV were 0.806, 0.600, 0.905, 0.750, and 0.826, respectively.
Conclusion
In conclusion, the combined DL model demonstrates the promising potential to predict SLM in patients with RC, thereby making individualized imaging test strategies.Acknowledgements
We thank all members of the author group for their efforts. We are also grateful for the support of all the patients and their families who participated in this work.References
1. Siegel, R.L., K.D. Miller, and A. Jemal, Cancer statistics, 2019. CA: A Cancer Journal for Clinicians, 2019. 69(1): p. 7-34.2. Keller, D.S., et al., The multidisciplinary management of rectal cancer. Nat Rev Gastroenterol Hepatol, 2020. 17(7): p. 414-429.
3. Van Cutsem, E., et al., Metastatic colorectal cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol, 2014. 25 Suppl 3: p. iii1-9.
4. Milette, S., et al., Molecular Pathways: Targeting the Microenvironment of Liver Metastases. Clin Cancer Res, 2017. 23(21): p. 6390-6399.
5. de Ridder, J.A.M., et al., Management of liver metastases in colorectal cancer patients: A retrospective case-control study of systemic therapy versus liver resection. Eur J Cancer, 2016. 59: p. 13-21.
6. Chow, F.C. and K.S. Chok, Colorectal liver metastases: An update on multidisciplinary approach. World J Hepatol, 2019. 11(2): p. 150-172.
7. Zhu, H.B., et al., Deep learning-assisted magnetic resonance imaging prediction of tumor response to chemotherapy in patients with colorectal liver metastases. Int J Cancer, 2021. 148(7): p. 1717-1730.
8. Liu, X., et al., Deep learning radiomics-based prediction of distant metastasis in patients with locally advanced rectal cancer after neoadjuvant chemoradiotherapy: A multicentre study. EBioMedicine, 2021. 69: p. 103442.
9. Wei, J., et al., Deep learning-based radiomics predicts response to chemotherapy in colorectal liver metastases. Med Phys, 2021. 48(1): p. 513-522.
10. He, K., et al., Deep Residual Learning for Image Recognition. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016: p. 770-778.
Figures