0714
Multiparametric radiomics-based machine learning predicts consensus molecular subtype 4 of colorectal cancer: a multi-center study1Fudan University Shanghai Cancer Center, Shanghai, China, 2ShanghaiTech University, Shanghai, China, 3MR Application Development, Siemens Shenzhen Magnetic Resonance Ltd, Shenzhen, China
Synopsis
Keywords: Pelvis, Multimodal
Motivation: The consensus molecular subtype (CMS) is a novel classification system that reflects the genetic characteristics of the tumor. CMS4 is associated with the worst prognosis.
Goal(s): To investigate whether a radiomics-based machine learning approach could predict CMS4 status in CRC patients.
Approach: The sequencing data was input into the CMS classification system to generate CMS subtype outcomes. Radiomics features were extracted from baseline T2WI and contrast-enhanced MRI. Machine learning algorithms were applied to explore the best-performing and most robust model.
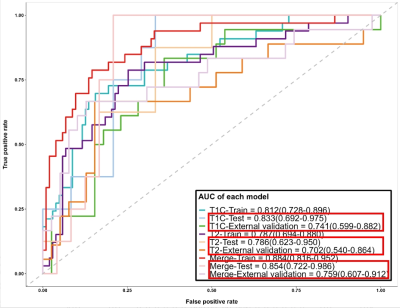
Results: The best performing model achieved AUCs of 0.855 and 0.759 in the test set and external validation set.
Impact: The genetic phenotype of CMS4 colorectal cancer may be potentially associated with morphological features. Multiparametric radiomics-based machine learning shows promising potential in distinguishing CMS4 from other subtypes of CRC.
INTRODUCTION
Gene expression profiles are widely recognized to be associated with tumor heterogeneity and therapeutic response in colorectal cancer (CRC). The consensus molecular subtype (CMS) is a novel classification system that reflects the genetic characteristics of the tumor. Among the four subtypes, CMS4 is associated with the worst prognosis. Patients with CMS4 are generally resistant to adjuvant chemotherapy treatment and have a higher likelihood of having micro metastases at the time of initial diagnosis due to its "early dissemination" pattern. Therefore, early detection of CMS4 is crucial for administering effective treatment protocols at an early stage. However, current CMS detection methods have several limitations, such as the insufficiency of biopsy samples and the destruction of tumor integrity after treatment, which hinder its early implementation. In CMS4 CRC, the upregulation of the EMT pathway and overexpression of TGF-β result in a histomorphology phenotype characterized by high tumor mesenchymal content and an angiogenesis-driven microenvironmental vascular abundance. These features may be reflected in MRI image features to some extent. Therefore, it is feasible to achieve early detection of CMS classification using an MRI-based radiomics approach. This study aimed to investigate whether a radiomics-based machine learning approach could predict CMS4 status in CRC patients.METHODS
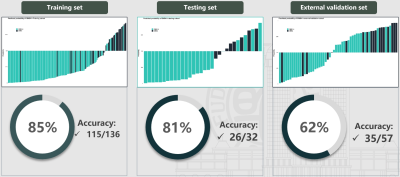
A total of 228 CRC cases from three centers were retrospectively included. Cases from center I were divided into training (138 cases) and validation sets (33 cases) in an 8:2 ratio; cases from center II and III were combined as the external validation set (57 cases). Sequencing data and Baseline MRI images, including T2-weighted (T2WI) and contrast-enhanced (CE) sequences, were available for each case. The sequencing data was input into the CMS classification system to generate CMS subtype outcomes. Radiomics features from the two sets were extracted with the same parameter settings. Several machine learning algorithms were applied in sample balance, feature normalization, feature filters, and classifier construction to explore the best-performing and most robust model for CMS4 prediction. The rad-score for each patient was calculated by the T2WI and CE models separately. The combined model was established by applying logistic regression on the results of the above two models. The performance of the model was evaluated using receiver operating characteristic (ROC) curve analysis. The area under the ROC curve (AUC) was calculated for quantification.RESULTS
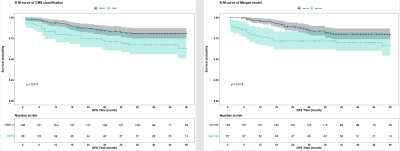
Among all cases, 59 (26%) cases were classified as CMS4. We found that the CE mode achieved better performance than T2 model in both the test set (0.815 vs 0.790) and external validation set (0.741 vs 0.702). After merging the two models, the predictive performance of the Merged model was further improved, with the AUCs of 0.855 and 0.759 in the test set and external validation set. The risk of recurrent metastasis in patients was effectively stratified by the the Merged model (P = 0.018).DISCUSSION
The genetic phenotype of CMS4 colorectal cancer may be potentially associated with morphological features. Multiparametric radiomics-based machine learning shows promising potential in distinguishing CMS4 from other subtypes of CRC.CONCLUSION
Multi-parametric imaging radiomics model developed based on CMS typing is expected to be a new non-invasive tool to effectively stratify the risk of recurrence in colorectal cancer patients, providing important information for personalized treatment plan development and adjustment.Acknowledgements
No acknowledgement found.References
No reference found.Figures