0713
Synthesized Gd-EOB-DTPA-enhanced hepatobiliary phase MR images via generative adversarial learning1Guangdong Cardiovascular Institute, Guangdong Provincial People's Hospital, Guangdong Academy of Medical Sciences, Guangzhou, China, GuangZhou, China, 2Department of Radiology, Guangdong Provincial People’s Hospital, Guangdong Academy of Medical Sciences, Southern Medical University, Guangzhou, China, Guangzhou, China, 3Guangdong Provincial Key Laboratory of Artificial Intelligence in Medical Image Analysis and Application, Guangdong Provincial People's Hospital, Guangdong Academy of Medical Sciences, Guangzhou, China, Guangzhou, China, 4Department of Medical Imaging Center, Nanfang Hospital, Southern Medical University, Guangzhou, China, Guangzhou, China, 5Philips Healthcare Guangzhou Ltd., Guangzhou, China
Synopsis
Keywords: Liver, Liver
Motivation: Gd-EOB-DTPA-enhanced hepatobiliary phase (HBP) imaging is clinical routine for liver lesion identification, and is usually empirically conducted at 20 minutes after bolus injection.
Goal(s): Our goal was to demonstrate the feasibility of optimizing clinical workflow by synthesizing Gd-EOB-DTPA-enhanced HBP images via machine learning.
Approach: Precontrast and early-enhanced T1WIs (5-min after bolus injection) acquired at 3 T were used to synthesize HBP images via a generative adversarial network in 490 subjects.
Results: Our preliminary results showed that synthesized HBP images are visually comparable to acquired HBP images with high SSIM(0.87±0.08) and PSNR(29.6±2.25).
Impact: Machine learning synthesized HBP images could provide comparable diagnostic information to acquired HBP images, suggesting that machine learning might be used to optimize clinical workflow and greatly shorten acquisition time for Gd-EOB-DTPA-enhanced MRI.
Introduction
Gadolinium ethoxybenzyl-diethylenetriamine pentaacetic acid (Gd-EOB-DTPA), a liver-specific contrast agent, can be specifically absorbed by hepatocytes and is widely adopted in dynamic contrast-enhanced MRI (DCE-MRI) for liver lesion identification (1). The peak absorption of Gd-EOB-DTPA could be obtained around 20 minutes after bolus injection, and T1WIs acquired at this phase are termed hepatobiliary phase (HBP) images (2). HBP images provide superior image quality, sensitivity, and specificity in detecting focal liver lesions, while the optimum delay time for an adequate HBP is controversial (3). Recent research suggested a personalized delay time based on patient’s liver function and the delay time for HBP might be reduced to 10 minutes in patients with normal liver function (4). The enhancement characteristics of liver tissues are supposed to be tightly related to the functionality of hepatocytes and the different functionalities between normal and dysfunctioned hepatocytes might be discriminated from early enhancement (5). Recognizing this opportunity, here we propose to adopt a machine-learning algorithm to learn the latent enhancement characteristics and synthesize HBP images from precontrast and early-enhanced T1WIs, i.e. 5-min after bolus injection, to greatly reduce the total scan time in clinical workflow.Method
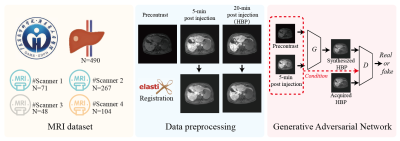
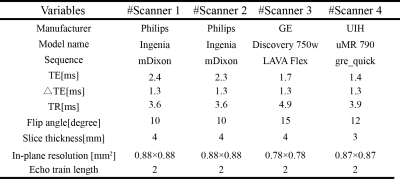
MRI DataThis retrospective study was approved by local institutional review aboard, and 490 Gd-EOB-DTPA-enhanced liver DCE-MRI scans that include precontrast, early-enhanced (5-min post bolus injection), and HBP (20-min post bolus injection) T1WIs were collected from 4 different scanners (3T). The details of the study design and MRI protocols are shown in Figure 1 and Figure 2, respectively.
Image preprocessing
All the T1WIs were first resampled to resolution of 1×1×3 mm3, and cropped or padded to matrix size of 384×384×64. To eliminate inter-phase motion, all the enhanced T1WIs were registered to precontrast T1WIs by using ITK-Elastix (6), and the registration performance was evaluated by using Dice coefficient and normalized mutual information (NMI) (7). The Dice coefficient was calculated in liver masks between precontrast and post-contrast T1WIs before and after volumetric registration, separately, where the liver masks were automatically segmented by a pretrained nnUNet model (8) and thereafter corrected by one experienced radiologist. After that, the T1WIs were normalized to signal intensity between -1 and 1.
Generative adversarial network
A conditional generative adversarial network of Pix2PixHD was trained for HBP synthesis on one Nvidia RTX 4090 GPU(24 GB) by using 80% randomly selected dataset and the remaining 20% dataset was used for testing (9). Pix2PixHD has been demonstrated superior performance in high-quality synthesis task with improved details and less blurring, by adopting a coarse-to-fine generator for synthesizing images at different resolutions, a multi-scale discriminator, and a feature-wise perceptual loss to mitigate unperfect registration-induced image blurring in synthesized images (10). During training, data augmentation including random flip, random rotation, random affine transform and random Gaussian noise(with standard deviation of 0.1) were adopted by using PyTorch.
Evaluation metrics
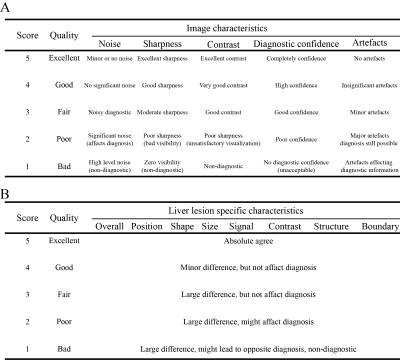
Quantitative metrics of structural similarity (SSIM) and peak signal-to-noise ratio (PSNR) were used for the objective evaluation of synthesized images. Qualitative evaluation of image quality and liver lesion-specific characteristics was blindly reviewed by one experienced radiologist (>15 years) in random order of synthesized HBP and acquired HBP images by using 5-point Likert Scale table as shown in Figure 3.
Results
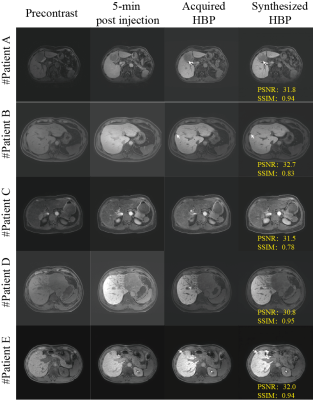
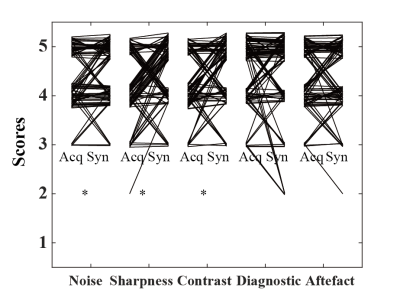
Image registration by ITK-Elastix significantly eliminates inter-phase motion with an improved Dice coefficient and NMI in 5-min post-contrast ( Dice: 0.87±0.10 vs. 0.95±0.04, p<0.001, NMI: 0.70±0.10 vs. 0.82±0.11, P<0.001) and in HBP( 0.85±0.11 vs. 0.95±0.03, p<0.001, NMI: 0.65±0.09 vs. 0.79±0.11, P<0.001). Five representative synthesized HBP images are shown in Figure 4. The synthesized HBP images are visually comparable to the acquired HBP images with high quantitative metrics of SSIM (0.87±0.08) and PSNR (29.6±2.25). The quanlitative evaluation showed slightly improvement in image characteristics of noise (4[IQR:4-5] vs. 5[IQR:4-5], P<0.05), sharpness (4[IQR:4-5] vs. 5[IQR:4-5], P<0.001), contrast (4[IQR:4-5] vs. 5[IQR:4-5], P<0.01), while there is no significant difference in diagnostic confidence (5[IQR:4-5] vs. 5[IQR:4-5], P=0.36) and artefacts (4[IQR:4-5] vs. 4.5[IQR:4-5], P=0.87). Liver lesion specific evaluation demonstrated excellent consistency within liver (5[IQR:4-5]) and lesion position (5[IQR:5-5]), and good consistency within signal(4[IQR:4-5]), size (4[IQR:4-4]), shape (4[IQR:4-4]), contrast(4[IQR:4-5]), structure (4[IQR:3-4]), and boundary (4[IQR:4-5]) of lesions, between synthesized HBP and acquired HBP images.Conclusion
Our preliminary study demonstrated synthesized HBP images could provide comparable diagnostic information to acquired HBP images, suggesting that synthesized HBP might be used to optimize the clinical workflow and greatly reduce total scan time in Gd-EOB-DTPA enhanced DCE-MRI. In addition, a comprehensive evaluation of diagnostic value of synthesized HBP images in liver lesion detection and characterization by radiologists is warranted in future studies.Acknowledgements
NoReferences
1. Jhaveri K, Cleary S, Audet P, et al. Consensus statements from a multidisciplinary expert panel on the utilization and application of a liver-specific MRI contrast agent (gadoxetic acid). AJR Am J Roentgenol 2015;204(3):498-509.
2. Hamm B, Staks T, Mühler A, et al. Phase I clinical evaluation of Gd-EOB-DTPA as a hepatobiliary MR contrast agent: safety, pharmacokinetics, and MR imaging. Radiology 1995;195(3):785-792.
3. Wang C, Yuan XD, Wu N, Sun WR, Tian Y. Optimization of hepatobiliary phase imaging in gadoxetic acid-enhanced magnetic resonance imaging: a narrative review. Quant Imaging Med Surg 2023;13(3):1972-1982.
4. Zech CJ, Ba-Ssalamah A, Berg T, et al. Consensus report from the 8th International Forum for Liver Magnetic Resonance Imaging. Eur Radiol 2020;30(1):370-382.
5. Schuhmann-Giampieri G, Schmitt-Willich H, Press WR, Negishi C, Weinmann HJ, Speck U. Preclinical evaluation of Gd-EOB-DTPA as a contrast agent in MR imaging of the hepatobiliary system. Radiology 1992;183(1):59-64.
6. Konstantinos Ntatsis, Niels Dekker, Viktor van der Valk, et al. itk-elastix: Medical image registration in Python. PROC OF THE 22nd PYTHON IN SCIENCE CONF (SCIPY 2023) 2023.
7. Luo X, Zhuang X. X-Metric: An N-Dimensional Information-Theoretic Framework for Groupwise Registration and Deep Combined Computing. IEEE Transactions on Pattern Analysis and Machine Intelligence 2023;45(07):9206-9224.
8. Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 2021;18(2):203-211.
9. Ting-Chun Wang, Ming-Yu Liu, Jun-Yan Zhu, Andrew Tao, Jan Kautz, Catanzaro B. High-Resolution Image Synthesis and Semantic Manipulation with Conditional GANs. CVPR 2018.
10. Johnson J, Alahi A, Fei-Fei L. Perceptual Losses for Real-Time Style Transfer and Super-Resolution. In: Leibe B, Matas J, Sebe N, Welling M, editors; 2016.
Figures