0341
Joint sequence optimization beats pure neural network approaches for super-resolution TSE1Institute of Neuroradiology, University Hospital Erlangen, Friedrich-Alexander University Erlangen-Nürnberg (FAU), Erlangen, Germany, 2Technical University of Munich, Munich, Germany, 3Munich Center for Machine Learning, Munich, Germany, 4Magnetic Resonance Center, Max-Planck-Institute for Biological Cybernetics, Tuebingen, Germany, 5Department of Computer Science, Friedrich-Alexander-Universität Erlangen-Nürnberg (FAU), Erlangen, Erlangen, Germany, 6Department Artificial Intelligence in Biomedical Engineering, Friedrich-Alexander-Universität Erlangen-Nürnberg (FAU), Erlangen, Germany
Synopsis
Keywords: AI/ML Image Reconstruction, Machine Learning/Artificial Intelligence, super-resolution, turbo-spin-echo, joint-optimization
Motivation: TSE flip angle trains can have a strong influence on the actual resolution of the acquired image and have consequently a considerable impact on the performance of a super-resolution task.
Goal(s): We demonstrate the advantage of end-to-end optimization of sequence and neural network parameter compared to pure network training approaches.
Approach: This MR-physics-informed training procedure jointly optimizes radiofrequency pulse trains of a PD- and T2-weighted TSE and subsequently applied CNN to predict corresponding PDw and T2w super-resolution TSE images.
Results: The method generalizes from simulation-based optimization to in vivo measurements and acquired super-resolution images show higher accuracy compared to pure network training approaches.
Impact: Acquired super-resolution image may improve evaluation of the data. Reduction of acquisition time compared to direct high-resolution acquisition leads to increase in patient comfort and minimization of motion artifacts.
Introduction
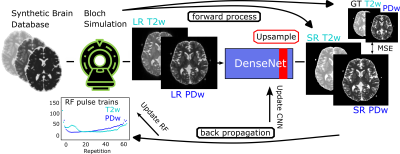
Current MRI super-resolution methods use contrasts acquired from typical clinical protocols as input for the neural network and disregard the influence of the MR sequence parameters for optimization.Using so-called known operator learning1, we propose an approach that utilizes a MR physics model during the optimization to not only train a neural network for super-resolution, but also adapt the refocusing RF pulses to directly influence the point-spread-function (PSF). This approach also allows the use of the uncorrupted theoretical contrast as ground truth, which is only available during the simulation. By using two different encoding schemes in our sequences, we gain additional information from the two different contrasts PD and T2 that are used as input for the CNN and both will have different PSFs, thus provide valuable information for the SR task. Both sequences are optimized jointly to allow generation of optimal contrasts for the SR task of the neural network.
Methods
A single-shot 2D TSE sequence is being used as default sequence for our optimization. The acquisition time for the single-shot 2D TSE is 0.76 s at 1.56 mm in-plane resolution. The refocusing RF pulses of the PDw TSE with TE=12 ms and T2w TSE with TE=96 ms were optimized jointly. All simulations and optimizations were performed in a fully differentiable Bloch simulation framework2.The forward simulation outputs the TSE signal which is conventionally reconstructed to magnitude images, and in addition the corresponding contrast as ground truth target. The entire process – MRI sequence, reconstruction, and evaluation – is modelled as one computational chain and is part of the forward and backwards propagation during the optimization, as depicted in Figure 1.After the optimization process, all sequences were exported using the Pulseq standard3 and the pypulseq tool4. Pulseq files could then be interpreted on a real MRI scanner including all necessary safety checks, and were executed on a 3T scanner using a 20-channel head coil. As high-resolution reference a vendor-provided TSE sequence was acquired with following parameters: 32-shot segmented, GRAPPA2, TE=12/96 ms, TR=12 s, FOV=200 mm×200 mm, matrix of 256×256, FA=180°. All MRI scans were performed after written informed consent was obtained.
Results
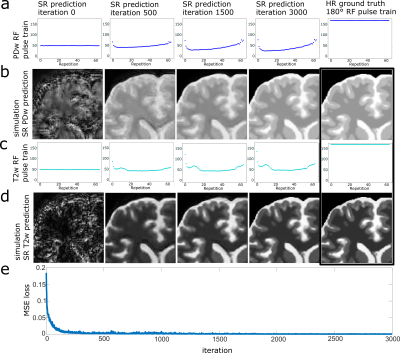
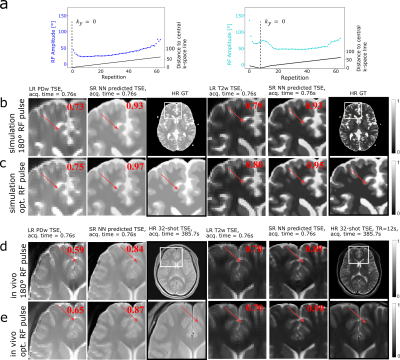
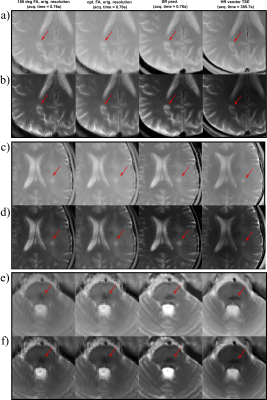
The optimization process can be seen in Figure 2. Starting from the initialized values, the RF pulses converge to the optimal RF pulse train, while the NN parameters are optimized simultaneously. The original LR TSE image with the zero-filled image and the reconstructed SR image are compared to our optimized RF pulse train design and a conventional 180° RF pulse train TSE sequence for each contrast in Figure 3. It can be observed, that in all cases the SR image leads to an improvement over the LR TSE image by showing clearer resolved borders between white and gray matter. The optimized RF pulse train further improves the nominal resolution, which can be observed by a clear increase of sharpness of the sulcus between Gyrus cinguli and Gyrus frontalis superior as indicated by the red arrows. The optimized sequence and CNN translate well to in vivo measurements, where similar improvements as seen in the simulated images can be observed (Figure 3d,e). Further, patient data have been acquired, three different pathologies with multiple sclerosis, glioblastoma and cavernoma. The PDw and T2w SR image of the patients are shown in Figure 4 with the red arrow indicating area of interest, respectively. The SR image demonstrates shaper edges of the lesion with clearer boundaries to the surrounding white matter.Discussion
We demonstrated a new end-to-end learning process for TSE super-resolution by jointly optimizing refocusing RF pulse trains and neural network parameters. This approach utilizes a differentiable MR physics simulation embedded in the forward and backward propagation. The joint-optimization outperforms a pure neural network training. Although our approach is solely based on simulated data, the optimized sequence and trained CNN translate well to in vivo data. Our approach is compatible with any network architecture e.g. 5-7 to further improve the SR task. By using simulation-based training data, we are able to use the theoretical uncorrupted contrast as ground truth target. Ground truth HR images measured in vivo are difficult to acquire due to the longer scan time. Motion artifacts become more significant and to acquire the same contrast as the LR counterpart the bandwidth has to be increased, leading to a decrease of SNR8.Conclusion
We propose an end-to-end optimization of MR sequence and neural network parameters for TSE super-resolution. This flexible and general end-to-end approach benefits from a MR physics informed training procedure, allowing a simple target-based problem formulation, and outperforms pure neural network training.Acknowledgements
No acknowledgement found.References
- Maier, A.K. et al.: Learning with known operators reduces maximum error bounds. Nature Machine Intelligence 2019 1:8. 1, 373–380 (2019).
- Loktyushin, A., et al. "MRzero‐Automated discovery of MRI sequences using supervised learning." Magn Reson Med 86.2 (2021): 709-724.
- Layton, K.J. et al.: Pulseq: A rapid and hardware-independent pulse sequence prototyping framework. Magn Reson Med. 77, 1544–1552 (2017).
- Ravi, Keerthi, Sairam Geethanath, and John Vaughan. "PyPulseq: A Python Package for MRI Pulse Sequence Design." Journal of Open Source Software 4.42 (2019): 1725.
- Li, G., Lyu, J., Wang, C., Dou, Q., Qin, J.: WavTrans: Synergizing Wavelet and Cross-Attention Transformer for Multi-contrast MRI Super-Resolution. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). 13436 LNCS, 463–473 (2022).
- Sui, Y., Afacan, O., Gholipour, A., Warfield, S.K.: MRI Super-Resolution Through Generative Degradation Learning. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). 12906 LNCS, 430–440 (2021).
- Lyu, Q., Shan, H., Wang, G.: MRI Super-Resolution with Ensemble Learning and Complementary Priors. IEEE Trans Comput Imaging. 6, 615–624 (2019).
- Portnoy, S., Kale, S.C., Feintuch, A., Tardif, C., Pike, G.B., Henkelman, R.M.: Information content of SNR/resolution trade-offs in three-dimensional magnetic resonance imaging. Med Phys. 36, 1442–1451 (2009).
Figures