1161
Super Resolution of MR Images with Deep Learning Based k-space Interpolation1Medical Image Processing Research Group (MIPRG), Dept. of Elect. & Comp. Engineering, COMSATS University, Islamabad, Pakistan
Synopsis
Super resolution of MR images can be used to speed up MRI scan time. However, super resolution is a highly ill-posed problem as the low-resolution images lack high frequency spatial information. In this paper, we propose a hybrid dual domain cascaded U-Net to restore the high-resolution images. Firstly, the U-Net operating in k-space domain is used to interpolate the missing k-space data points and then the U-Net operating in image domain provides a refined high-resolution solution image. Experimental results show a successful reconstruction of high resolution images by using only central 6.25% and 25% k-space data.
Introduction
Magnetic resonance imaging (MRI) is widely used in the detection and diagnosis of diseases. High-resolution MR images help clinicians to locate lesions and diagnose diseases. However, the acquisition of high-resolution MR images requires high magnetic field intensity and long scanning time, which will bring discomfort to patients and easily introduce motion artifacts, resulting in image quality degradation1. Super resolution is a technique to restore high-frequency details from a low-resolution image to improve image resolution3. The simplest method of super resolution could be an interpolation technique3, such as bicubic interpolation7 or nearest neighbor interpolation2. However, the interpolation method does not, in essence, increase the image information, so it cannot recover the high-frequency information of an image2. In this paper, we propose a hybrid dual domain cascaded deep learning framework for generating high resolution images from the corresponding low resolution k-space data.Method
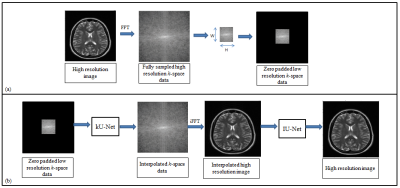
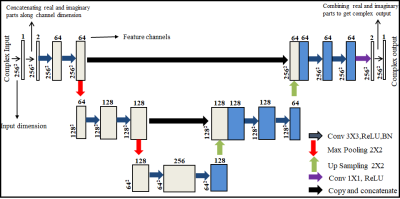
The fundamental goal of super resolution is to collect the missing information by reconstructing the super-resolved solution images from the low-resolution input images. The low-resolution images miss the high spatial frequency information which is provided by the periphery of the k-space data4. In the proposed deep learning framework (Figure 1), first the customized architecture of U-Net6 is used in the k-space domain (denoted as kU-Net) to interpolate the missing k-space information in the low-resolution data. The kU-Net estimates the interpolated image which is later refined by the customized architecture of U-Net in image domain (denoted as IU-Net). For the proposed method, the training dataset is extracted from 1407 human brain Cartesian dataset (of 30 Multiple Sclerosis patients) acquired from 1.5T scanner5. These MR images were obtained using a T2-weighted turbo spin echo pulse sequence.The original high-resolution images (of size (r,c), where r is the number of rows and c is the number of columns) are converted into fully sampled k-space data via 2D Fourier transform. Next, low-resolution k-space data is generated by windowing a central region (of size (W,H), where W is the width and H is the height of the window) of the fully sampled k-space data. In this way, the MX fold resolution degradation (also called Down Sampling Factor (DSF)) is defined as r/W and c/H. The zero-padded low-resolution k-space data is used as an input whereas the corresponding fully sampled k-space data is used as the ground truth for the training of kU-Net. The k-space data is complex-valued in nature, hence for the training of kU-Net, the real and imaginary parts of the complex k-space data are concatenated along the channel dimension.
The output of the trained kU-Net is the interpolated complex k-space data. IFFT of the interpolated complex-valued k-space data provides an interpolated image which is given as an input to the IU-Net for further processing.
The IU-Net is trained in the image domain to refine the coarse interpolated images. For this purpose, the output of kU-Net is used as an input whereas fully sampled high-resolution images are used as a ground truth for the training of IU-Net. The real and imaginary parts of the complex-valued MR images are concatenated along the channel dimension for the training of IU-Net.
In our proposed method, kU-Net and IU-Net are trained independently. For training of kU-Net and IU-Net, all the weights of the convolutional layers were initialized by a zero-centered normal distribution with standard deviation of 0.05 without a bias term. The loss function was minimized by using the RMSPropOptimizer with a learning rate of 0.001, mini-batch size of 2, and 1000 epochs. Training was implemented on Python 3.7.1 by Keras using TensorFlow as a backend on Intel(R) core (TM) i7-4790 CPU, clock frequency 3.6GHz, 16 GB RAM and GPU NVIDIA GeForce GTX 780 for approximately eighteen hours. The proposed method was tested on 431 low-resolution human head data5. The high-resolution results obtained with the proposed method are compared against bicubic interpolation7.
Results
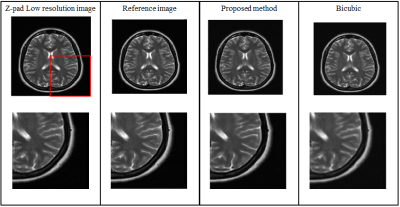
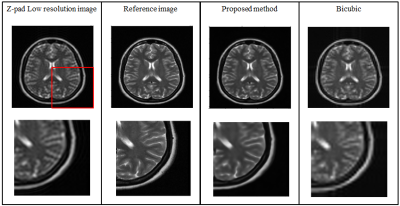
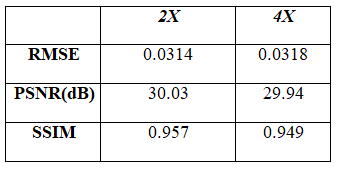
Figure 3 and 4 show the super resolution results obtained from the proposed method for DSF of 2X and 4X, respectively. Table 1 shows the Root Mean Square Error (RMSE), Peak Signal-to-Noise Ratio (PSNR) and Structural Similarity Index (SSIM) values of the super resolution results obtained from the proposed method for DSF of 2X and 4X.Discussion and Conclusion
The low-resolution images obtained with DSF of 2X and 4X retain only central 25% and 6.25% k-space data, respectively. Hence, with the help of the proposed method, we have successfully reconstructed images with the central 6.25% and 25% k-space data, respectively. Therefore, the proposed super resolution technique can help to reduce the scan time and avoid motion artifacts especially in cardiac imaging. Compared to conventional super resolution technique (i.e. bicubic interpolation7), the super resolution results obtained from the proposed method are more sharp and close to reference image as indicated by their PSNR, SSIM and RMSE values.Acknowledgements
No acknowledgement found.References
1. Liu, Huanyu, et al. "DL-MRI: A Unified Framework of Deep Learning-Based MRI Super Resolution." Journal of Healthcare Engineering 2021 (2021).
2. Masutani, Evan M., Naeim Bahrami, and Albert Hsiao. "Deep learning single-frame and multiframe super-resolution for cardiac MRI." Radiology 295.3 (2020): 552-561.
3. Li, Xiaofang, et al. "Deep learning methods in real-time image super-resolution: a survey." Journal of Real-Time Image Processing 17.6 (2020): 1885-1909.
4. D.W. McRobbie, E.A. Moore, M.J. Graves, MRI from picture to proton, 2017. https://doi.org/10.2214/ajr.182.3.1820592.
5. C.M. Hyun, H.P. Kim, S.M. Lee, S. Lee, J.K. Seo, Deep learning for undersampled MRI reconstruction, Phys. Med. Biol. 63 (2018) aac71a. https://doi.org/10.1088/1361-6560/aac71a.
6. Ronneberger, Olaf, Philipp Fischer, and Thomas Brox. "U-net: Convolutional networks for biomedical image segmentation." International Conference on Medical image computing and computer-assisted intervention. Springer, Cham, 2015.
7. Ashikaga, Hiroshi, et al. "Quantitative assessment of single-image super-resolution in myocardial scar imaging." IEEE journal of translational engineering in health and medicine 2 (2014): 1-12.
Figures