2669
Ultra-Fast Simultaneous T1rho and T2 Mapping Using Deep Learning1Department of Biomedical Engineering, Department of Electrical Engineering, The State University of New York at Buffalo, Buffalo, NY, United States, 2Program of Advanced Musculoskeletal Imaging (PAMI), Cleveland Clinic, Cleveland, OH, United States, 3Paul C. Lauterbur Research Center for Biomedical Imaging, Medical AI research center, SIAT, CAS, Shenzhen, China
Synopsis
This abstract presents a deep learning method to generate T1rho and T2 relaxation maps simultaneously within one scan. The method uses 3D deep convolutional neural networks to exploit the nonlinear relationship between and within the combined subsampled T1rho and T2-weighted images and the combined T1rho and T2 maps, bypassing conventional fitting models. Compare with separated trained relaxation maps, this new method also exploits the autocorrelation and cross-correlation between subsampled echoes. Experiments show that the proposed method is capable of generating T1rho and T2 maps simultaneously from only 3 subsampled echoes within one scan with quantification results comparable to reference maps.
Introduction
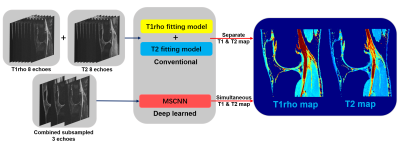
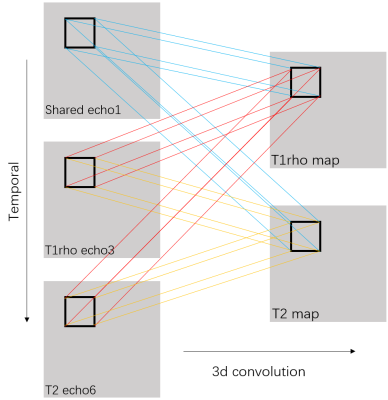
T1rho and T2 imaging have gained much attention as quantitative MRI methods to detect cartilage matrix changes; with T2 measures being sensitive to hydration and collagen structure and contents and T1rho measures being more sensitive to proteoglycan contents1. They are promising imaging biomarkers for osteoarthritis (OA) that can be potentially used in large multicenter clinical trials and translated into clinical practice. However, 4-8 echoes are typically necessary for reliable estimation of a parameter map in the conventional method. Such a long acquisition time prevents parameter mapping from being widely used clinically. It is of great interest to accelerate quantitative imaging. Deep learning methods have been used recently to accelerate MR acquisition by reconstructing images from subsampled k-space data2-7. The learning-based reconstruction methods have the benefit of ultrafast online reconstruction once the offline training is completed. Although there are quite some works on deep reconstruction of morphological images, very few works have studied tissue parameter mapping8-10. As shown in figure 1, we develop a deep learning-based framework for ultrafast quantitative MR imaging with multi-channel input (images from different echoes) and multi-channel output (different parameter maps) based on our previous network11 for diffusion tensor imaging. The purpose of this study is to demonstrate the feasibility of such framework, called Model Skipped Convolutional Neural Network (MSCNN), for ultrafast simultaneous T1rho and T2 mapping of the knee within one scan. Using 3D convolutional kernels as shown in figure 2, we demonstrate for the first time the feasibility to simultaneously generate T1rho and T2 maps using as few as 3 echoes, where echo 1 is a shared echo with both T1rho and T2 weighting, echo 3 is a T1rho-weighted image, and echo 6 is a T2-weighted image. These 3 echoes are further subsampled in k-space to provide additional acceleration. Collecting T1rho- and T2-weighted images in one acquisition using the exact same readout will facilitate comparisons of these two measures and allow exploration of interesting new markers such as R2-R1rho.Theory and Methods
In line with our prior study for diffusion tensor imaging using reduced acquisition to generated multi-channel colormap11, the loss term $$$L(Θ)=\frac{1}{n}\sum_{i=1}^{n}\| F(x_{i};Θ) -y_{i}\|^{2} (1)$$$ ensures that the reconstructed parameter maps from end-to-end CNN mapping are accurate compared to the maps from fully sampled echoes while bypassing the conventional error-prone model fitting step. Network parameters $$$Θ$$$ is MSE-base learned that minimize the multi-channel loss function. For our 3D convolutional neural network, ten weighted layers are used for training and testing. For each layer except the last one, 64 filters with kernel size of 3 are used. Specifically, we use a weight decay of 0.0001 and a learning rate of 0.0001 for 1.4M iterations with ReLU as the activation function (except for the output layer). In the testing stage, the acquired 3 echoes are fed into the CNN with learned $$$Θ$$$ to generate the desired quantitative maps $$$F(x_t;Θ)$$$.Ten sets of knee data were collected at a 3T MR scanner (Prisma, Siemens Healthineers) with a 1Tx/15Rx knee coil (QED), using a magnetization-prepared angle-modulated partitioned k-space spoiled gradient echo snapshots (MAPSS) T1ρ and T2 quantification sequences (time of spin-lock [TSLs] of 0, 10, 20, 30, 40, 50, 60, 70ms, spin-lock frequency 500Hz, Preparation TEs of 0, 9.7, 21.3, 32.9, 44.5, 56.1, 67.6, 79.2 ms, matrix size 160×320×8×24 [PE×FE×Echo×Slice], FOV 14cm, and slice thickness 4mm). Among these data, 8 sets were used to train the proposed MSCNN and 2 for testing. For echo subsampling, the 1st, 3rd and 6th echoes are selected. For images within selected echoes, 2D Poisson random sampling was used with an acceleration factor of 2 for further reduction. Hardware specification: i9 7980XE; 64 GB; GPU 2x NVIDIA Quadro P6000. The training takes around 15 hours. It takes only 0.08 seconds to generate a complete set of combined T1rho and T2 maps using the learned network while it takes around 30 min to get both maps using conventional exponential decay fitting.
Results
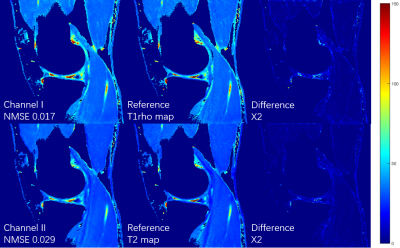
Figures 3 shows the T1rho maps and T2 maps, respectively, from a total of 3 echoes generated using the proposed deep learning method. Poisson disk undersampling with a reduction factor of 2 is used in k-space subsampling within these 3 echoes. Results from all 8 echoes using conventional fitting model were used as the ground truth for comparison. The combined reduction factor is 10.66. (2 x 8 echoes/3 selected-echoes x RF2 in k-space) It can be seen that the quantitative maps generated by our deep learning method are very close to the ground truth even with only 3 subsampled echoes. The result is further verified by the normalized mean squared errors (NMSE) shown on the bottom left of each image.Conclusion
In this abstract, we presented a deep convolutional neural network called MSCNN for superfast MR quantitative imaging. Experimental results showed that our proposed network is capable of generating T1rho and T2 maps from only 3 subsampled echo images within one scan. With a scan time of only 5 min, we were able to obtain a complete set of joint T1rho and T2 maps. Comparison of combined and separated training using larger datasets will be performed for evaluation of quantification accuracy and diagnostic performance.Acknowledgements
The work was partly supported by the Arthritis Foundation.References
[1] Link TM, Neumann J, Li X. Prestructural cartilage assessment using MRI. J Magn Reson Imaging. 2017 Apr;45(4):949-965. doi: 10.1002/jmri.25554. Epub 2016 Dec 26.
[2] Wang S, Su Z, Ying L, Peng X, Liang D, et al. Accelerating magnetic resonance imaging via deep learning. IEEE 14th International Symposium on Biomedical Imaging (ISBI), pp. 514-517, Apr. 2016.
[3] Wang S, Huang N, Ying L, Liang D, et al. 1D Partial Fourier Parallel MR imaging with deep convolutional neural network. Proceedings of International Society of Magnetic Resonance in Medicine Scientific Meeting, 2017.
[4] Lee D, Yoo J, et al. Deep residual learning for compressed sensing MRI. IEEE 14th International Symposium on Biomedical Imaging (ISBI), Melbourne, VIC, pp. 15-18, Apr. 2017.
[5] Wang S, Zhao T, Ying L, and Liang D. Feasibility of Multi-contrast MR imaging via deep learning. Proceedings of International Society of Magnetic Resonance in Medicine Scientific Meeting, 2017.
[6] Schlemper J, Caballero J, et al. A Deep Cascade of Convolutional Neural Networks for Dynamic MR Image Reconstruction. arXiv:1704.02422, 2017.
[7] Jin K H, McCann M T et al. Deep Convolutional Neural Network for Inverse Problems in Imaging. in IEEE Transactions on Image Processing, vol. 26, no. 9, pp. 4509-4522, Sept. 2017.
[8] Liu F, Feng L, Kijowski R. MANTIS: Model-Augmented Neural neTwork with Incoherent k-space Sampling for efficient estimation of MR parameters. International Society of Magnetic Resonance in Medicine Machine Learning Workshop Part II, 2018.
[9] Zhang Q, Su P, Liao Y, et al. Deep learning based MR fingerprinting ASL ReconStruction. Proceedings of 27th ISMRM Annual Meeting, Montreal, Canada, #0820, 2019.
[10] Pirkl C, Lipp I, Buonincontri G, et al. Deep Learning-enabled Diffusion Tensor MR Fingerprinting. Proceedings of 27th ISMRM Annual Meeting, Montreal, Canada, #1102, 2019.
[11] Li H, Zhang C, Liang Z, Liang D, Shen B, et al. Deep Learned Diffusion Tensor Imaging. Proceedings of 27th ISMRM Annual Meeting, Montreal, Canada, #3344, 2019.
Figures