0993
RED-N2N: Image reconstruction for MRI using deep CNN priors trained without ground truth1Washington University in St. Louis, St. Louis, MO, United States
Synopsis
We propose a new MR image reconstruction method that systematically enforces data consistency while also exploiting deep-learning imaging priors. The prior is specified through a convolutional neural network (CNN) trained to remove undersampling artifacts from MR images without any artifact-free ground truth. The results on reconstructing free-breathing MRI data into ten respiratory phases show that the method can form high-quality 4D images from severely undersampled measurements corresponding to acquisitions of about 1 minute in length. The results also highlight the improved performance of the method compared to several popular alternatives, including compressive sensing and UNet3D.
Introduction
The problem of forming an image from undersampled k-space measurements is common in MRI. For example, free-breathing MRI uses self-navigation techniques for detecting respiratory motion from data, which is subsequently binned into multiple respiratory phases, thus resulting in sets of undersampled k-space measurements [1–4]. To mitigate streaking artifacts due to undersampling, it is common to use an imaging prior, such as transform-domain sparsity [5–7]. More recently, deep learning has been explored for overcoming undersampling artifacts in MRI [8–10].Regularization by denoising (RED) is a recent framework that specifies the imaging prior through an image denoiser [11]. RED explicitly accounts for the forward model in an iterative manner while exploiting powerful denoising convolutional neural networks (CNNs) as priors [12–16]. The CNN training requires a dataset containing matched pairs of noisy and noiseless ground-truth images, limiting the practical applicability. Recently, a new technique called Noise2Noise was introduced [17] for training CNNs given only pairs of noisy images.
We introduce a new RED algorithm that replaces a denoising CNN with a more general image restoration CNN. We observe that respiratory binning leads to different k-space coverage patterns for different acquisition times, leading to distinct artifact patterns. Based on this observation, and inspired by Noise2Noise, we learn our prior by mapping pairs of complex MR volumes acquired over different acquisition times to one another, without using artifact-free ground-truth images. The trained CNN is then introduced into the iterative RED algorithm, where it is combined with the k-space data consistency term. We refer to our technique as RED-N2N and apply it for acquisition times ranging from 1 to 5 minutes.
Methods
All experiments were performed on a 3T PET/MRI scanner (Biograph mMR; Siemens Healthcare, Erlangen, Germany). The data was collected using CAPTURE, a recently proposed T1-weighted stack-of-stars 3D spoiled gradient-echo sequence with fat suppression that has consistently acquired projections for respiratory motion detection [4]. Upon the approval of our Institutional Review Board, 15 healthy volunteers and 17 cancer patients were recruited.The acquisition parameters were as follows: TE/TR = 1.69 ms/3.54 ms, FOV = 360 mm x 360 mm, in-plane resolution = 1.125x1.125 mm, partial Fourier factor = 6/8, number of radial lines = 2000, slice resolution = 50%, slices per slab = 96 with a slice thickness of 3 mm, total acquisition time = about 5 minutes (slightly longer for larger subjects).
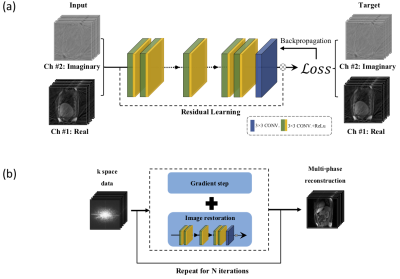
RED-N2N replaces the denoiser in RED by a 3D DnCNN network (x-y-phase) [18] trained for removing streaking artifacts from complex-valued MR volumes. The training of DnCNN was inspired by Noise2Noise and uses pairs of MR volumes corresponding to the same person, but acquired over different acquisition times with no ground truth data. Figure 1 exhibits the details of the RED-N2N method. We used 8 healthy subjects for training and 1 for validation. The remaining 6 healthy subjects and the 17 patients were used for testing. 400, 800, 1200 and 1600 radial spokes were used to reconstruct the images.
We evaluated the performance of RED-N2N against multi-coil non-uniform inverse fast Fourier transform (MCNUFFT), compressed sensing (CS) [4], and UNet3D (x-y-phase) trained by using the 5-minute CS reconstruction as the ground truth.
Results
Figure 3 and 4 show reconstructions for 400 radial spokes for two patients. The 2000-spoke CS serves as the gold-standard reference. The 400-spoke CS could not remove all artifacts, while Unet3D leads to some blurring in the reconstructed image. RED-N2N provided sharper reconstructions when compared to both methods. Figures 5 and 6 show Unet3D and RED-N2N reconstructions for 400, 800, 1200 and 1600 radial spokes for two patients. There is a noticeable improvement in quality for 800 spokes when compared to 400 spokes for both methods, but the quality remains relatively stable after 800 spokes.Discussion
RED-N2N reconstructs high-quality images even for 1-minute acquisitions. The performance of RED-N2N relative to CS and UNet3D indicates that by accounting for the forward model and using a learned prior, one can obtain significant improvements over the traditional model-based and learning-based methods. Remarkably, the performance of RED-N2N using the same DnCNN network – trained only on healthy subjects – generalizes to the data from patients that have significantly different image features.Conclusion
We proposed RED-N2N as an iterative method that combines a learned CNN prior with the data consistency in the undersampled k-space domain. RED-N2N offers best of model-based and learning-based worlds by outperforming CS and UNet3D, even without learning on artifact-free ground-truth data.Acknowledgements
No acknowledgement found.References
1. Grimm R, Fürst S, Souvatzoglou M, et al. Self-gated MRI motion modeling for respiratory motion compensation in integrated PET/MRI. Med. Image Anal. 2015;19(1):110–120.
2. Feng, L, Grimm R, Block KT, et al. Golden-angle radial sparse parallel MRI: combination of compressed sensing, parallel imaging, and golden-angle radial sampling for fast and flexible dynamic volumetric MRI. Magn. Reson. Med. 2014;72(3):707–717.
3. Feng L, Axel L, Chandarana H, et al. XD-GRASP: Golden-angle radial MRI with reconstruction of extra motion-state dimensions using compressed sensing. Magn. Reson. Med. 2016;75(2):775–788.
4. Eldeniz C, Fraum T, Salter A, et al. CAPTURE: Consistently Acquired Projections for Tuned and Robust Estimation: A Self-Navigated Respiratory Motion Correction Approach. Invest Radiol. 2018;53(5):293–305.
5. Lustig M, Donoho DL, Pauly JM. Sparse MRI: The Application of Compressed Sensing for Rapid MR Imaging. Magn. Reson. Med. 2007;58(6):1182–1195.
6. Knoll F, Brendies K, Pock T, et al. Second Order Total Generalized Variation (TGV) for MRI. Magn. Reson. Med. 2011;65(2):480–491.
7. Otazo R, Candès E, Sodickson DK. Low-Rank Plus Sparse Matrix Decomposition for Accelerated Dynamic MRI with Separation of Background and Dynamic Components. Magn. Reson. Med. 2015;73:1125–1136.
8. Han YS, Yoo J, Ye JC. Deep learning with domain adaptation for accelerated projection‐reconstruction MR. Magn. Reson. Med. 2017;80(3):1189–1205.
9. Lee D, Yoo J, Tak S, et al. Deep Residual Learning for Accelerated MRI Using Magnitude and Phase Networks. IEEE Trans. Biomed. Eng. 2018;65(9):1985–1995.
10. Aggarwal HK, Mani MP, Jacob M. MoDL: Model Based Deep Learning Architecture for Inverse Problems. IEEE Trans. Med. Imag. 2018;38(2):394–405.
11. Romano Y, Elad M, Milanfar P. The Little Engine That Could: Regularization by Denoising (RED). SIAM J. Imaging Sci. 2017;10(4):1804–1844.
12. Reehorst ET, Schniter P. Regularization by Denoising: Clarifications and New Interpretations. IEEE Trans. Comput. Imag. 2019;5(1):52–67.
13. Metzler CA, Schniter P, Veeraraghavan A, et al. prDeep: Robust Phase Retrieval with a Flexible Deep Network. In: Proc. 35th Int. Conf. Machine Learning (ICML). Stockholm, Sweden; 2018.
14. Sun Y, Liu J, Kamilov US. Block Coordinate Regularization by Denoising. In: Proc. Advances in Neural Information Processing Systems 33. Vancouver, BC, Canada; 2019.
15. Mataev G, Elad M, Milanfar P. DeepRED: Deep Image Prior Powered by RED. Proc. IEEE Int. Conf. Comp. Vis. Workshops (ICCVW). 2019.
16. Wu Z, Sun Y, Liu J, et al. Online Regularization by Denoising with Applications to Phase Retrieval. In: Proc. IEEE Int. Conf. Comp. Vis. Workshops (ICCVW). Seoul, Korea Republic; 2019.
17. Lehtinen J, Munkberg J, Hasselgren J, et al. Noise2Noise: Learning Image Restoration without Clean Data. In: Proc. 35th Int. Conf. Machine Learning (ICML). Stockholm, Sweden; 2018.
18. Zhang K, Zuo W, Chen Y, et al. Beyond a Gaussian Denoiser: Residual Learning of Deep CNN for Image Denoising. IEEE Trans. Image Process. 2017;26(7):3142–3155.
Figures