0991
Deep Learning MRI Reconstruction in Application to Point-of-Care MRI
Jo Schlemper1, Seyed Sadegh Mohseni Salehi1, Carole Lazarus1, Hadrien Dyvorne1, Rafael O'Halloran1, Nicholas de Zwart1, Laura Sacolick1, Samantha By1, Joel M. Stein2, Daniel Rueckert3, Michal Sofka1, and Prantik Kundu1,4
1Hyperfine Research Inc., Guilford, CT, United States, 2Hospitals of the University of Pennsylvania, Philadelphia, PA, United States, 3Computing, Imperial College London, London, United Kingdom, 4Icahn School of Medicine at Mount Sinai, New York City, NY, United States
1Hyperfine Research Inc., Guilford, CT, United States, 2Hospitals of the University of Pennsylvania, Philadelphia, PA, United States, 3Computing, Imperial College London, London, United Kingdom, 4Icahn School of Medicine at Mount Sinai, New York City, NY, United States
Synopsis
The goal of low-field (64 mT) portable point-of-care (POC) MRI is to produce low cost, clinically acceptable MR images in reasonable scan times. However, non-ideal MRI behaviors make the image quality susceptible to artifacts from system imperfections and undersampling. In this work, a deep learning approach is proposed for fast reconstruction from hardware and sampling-associated imaging artifacts. The proposed approach outperforms the reference deep learning approaches for retrospectively undersampled data with simulated system imperfections. Furthermore, we demonstrate that it yields better image quality and faster reconstruction than compressed sensing approach for unseen, prospectively undersampled low-field POC MR images.
Background
The goal of portable point-of-care (POC) MRI is to propose a low cost MRI that provides mobility, stability, and clinically acceptable images in reasonable scan times. Low-field (64 mT) POC MRI gives recognizable contrast with no ionizing radiation and little heating risk, however, the image quality is susceptible to artifacts from system imperfections, patient motion and undersampling. Standard MRI reconstruction of POC MRI data based on fast linear methods yield images that are clinically acceptable1-4, yet the reconstructed images still exhibit non-ideal imaging behaviors such as encoding error. If these errors are modeled accurately, they may be recovered by iterative reconstruction such as parallel imaging5-9 and compressed sensing10-16 (CS). Nevertheless, iterative methods have high computational cost and at low-field, high-level of noise and system imperfections further complicates the parameter tuning process. Inspired by the recent success of deep learning (DL) in MRI reconstruction17-25, we propose a DL approach that could perform simultaneous reconstruction and recovery from hardware and sampling-associated imaging artifacts. We postulate that such a reconstruction system could give higher quality POC MRI images with fast reconstruction time.Materials and Methods
Imaging: Research was performed with patient consent under an Institutional Review Board approved Study at Hospitals of the University of Pennsylvania. Imaging was done using the Hyperfine POC MRI (64 mT, 28 T/m/s slew rate, 15mT/m gradient, 8ch receive array, head-only volume transmit coil). A Fast Spin Echo (FSE) with 3D Cartesian encoding with variable Poisson disk (VPD) sampling26 in both phase encoding directions was used to acquire a 64-echo segmented into 8 images, which are combined subsequently. The field-of-view (FOV) is 220x180x180mm and the encoding resolution is 1.5x1.5x5mm. FLAIR and T2-weighted whole-brain images were acquired from 10 patients, where the scan times were 11:11s and 5:28s respectively.Training Data: Ground truth images consisted of 160 randomly selected T1- and T2-weighted MR volumes released by the Human Connectome Project27. To train a model that is robust to system imperfections, we generated training data of pairs of ground truth images and corresponding corrupted sensor data. This involved: 1) resampling the ground truth image to match Hyperfine T2-weighted FSE image FOV and resolutions, 2) adding image phase and coil sensitivity weighting, 3) apply non-uniform Fourier transform3 using the VPD sampling with random jittering of $$$k$$$-space points. Such degradation processes model a multichannel dataset acquired in the presence of eddy currents, interference and patient motion.
Model Architecture: We implemented a 2D Nonuniform Variational Network (NVN)28 for reconstructing the undersampled low-field MR images with the associated hardware-related systematic errors. NVN, as shown in Figure 1, is a technique based on the unrolling of nonlinear iterative reconstruction algorithms such as CS. It consists of multiple blocks representing the update steps in CS that are made learnable. Each block-$$$i$$$ has three components: CNN-$$$i$$$, DC-$$$i$$$ and $$$\lambda$$$. CNN-$$$i$$$ is a small network inspired by U-net29 for image de-aliasing. DC-$$$i$$$ is a data consistency layer30 extended for arbitrary, nonuniform $$$k$$$-space data, which ensures the reconstructed images to have high data fidelity. Finally, $$$\lambda$$$ enables the trade-off between de-aliasing and data consistency. The model has 6 blocks and is end-to-end trainable.
Evaluation: The proposed approach was first evaluated using 40 MR volumes from HCP data for quantitative validation using the metrics including MSE, SSIM31, and PSNR. NVN was compared to the U-net approaches24,29 for the simulated acquisition with acceleration factors 3.5. To evaluate the model generalizability to low-field MR image obtained with system imperfections, the proposed method was evaluated using prospectively undersampled FLAIR and T2-weighted images from Hyperfine MR device and compared against gridding1 and a wavelet-based CS-SENSE 3D reconstruction32,33.
Results
Results of the validation experiment are summarized in Figure 2 and sample reconstructions are shown in Figure 3. NVN achieved the best results quantitatively and resulted in the sharper reconstructions with the highest data fidelity. In Figure 4, the reconstruction from prospectively undersampled low-field MR data is shown. In particular, visible blurriness in gridding is mitigated in NVN and CS. The reconstruction times were 0.3s, 5.5s and 50s per 3D volume for gridding, NVN and CS-SENSE respectively.Discussion
Quantitative and qualitative analyses show the high capacity of NVN to learn a reconstruction process that is robust across different high-field MRI contrasts (T1, T2), low-field MRI contrasts (T2, FLAIR) and variable unknown noise levels. Note that the training data only consisted of high-field MRI contrasts and did not represent MRI contrast variation associated with 64 mT34. Yet, NVN generalized well. NVN primarily mitigated noise and blurriness attributed to encoding errors and undersampling. We associated such observed robustness to the NVN architecture (DC layer) and training with encoding errors. Further study is required on the bounds of this robustness. NVN also has fast reconstruction speed, making the proposed strategy well-suited to just-in-time reconstruction as needed in the POC MRI application.Conclusion
We established that a DL approach to POC MRI reconstruction could perform both reconstruction and error recovery in low computational time. We found a successful strategy in a learnable unrolled iterative reconstruction with data consistency blocks, where the importance was demonstrated by the robustness to the simulated HCP data and the prospectively undersampled low-field images.Acknowledgements
This work was supported by research funding from Hyperfine Research, Inc.References
- Zwart, Nicholas R., Kenneth O. Johnson, and James G. Pipe. "Efficient sample density estimation by combining gridding and an optimized kernel." Magnetic resonance in medicine 67.3 (2012): 701-710.
- Beatty, Philip J., Dwight G. Nishimura, and John M. Pauly. "Rapid gridding reconstruction with a minimal oversampling ratio." IEEE transactions on medical imaging 24.6 (2005): 799-808.
- Fessler, Jeffrey A. "On NUFFT-based gridding for non-Cartesian MRI." Journal of Magnetic Resonance 188.2 (2007): 191-195.
- Block, Kai Tobias, Martin Uecker, and Jens Frahm. "Undersampled radial MRI with multiple coils. Iterative image reconstruction using a total variation constraint." Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 57.6 (2007): 1086-1098.
- Sodickson, Daniel K., and Warren J. Manning. "Simultaneous acquisition of spatial harmonics (SMASH): fast imaging with radiofrequency coil arrays." Magnetic resonance in medicine 38.4 (1997): 591-603.
- Griswold, Mark A., et al. "Generalized autocalibrating partially parallel acquisitions (GRAPPA)." Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 47.6 (2002): 1202-1210.
- Pruessmann, Klaas P., et al. "SENSE: sensitivity encoding for fast MRI." Magnetic resonance in medicine 42.5 (1999): 952-962.
- Pruessmann, Klaas P., et al. "Advances in sensitivity encoding with arbitrary k‐space trajectories." Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 46.4 (2001): 638-651.
- Uecker, Martin, et al. "ESPIRiT—an eigenvalue approach to autocalibrating parallel MRI: where SENSE meets GRAPPA." Magnetic resonance in medicine 71.3 (2014): 990-1001.
- Lustig, Michael, David Donoho, and John M. Pauly. "Sparse MRI: The application of compressed sensing for rapid MR imaging." Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 58.6 (2007): 1182-1195.
- Murphy, Mark, et al. "Fast l1-SPIRiT Compressed Sensing Parallel Imaging MRI: Scalable Parallel Implementation and Clinically Feasible Runtime." IEEE transactions on medical imaging 31.6 (2012): 1250-1262.
- Knoll, Florian, et al. "Second order total generalized variation (TGV) for MRI." Magnetic resonance in medicine 65.2 (2011): 480-491.
- Ravishankar, Saiprasad, and Yoram Bresler. "MR image reconstruction from highly undersampled k-space data by dictionary learning." IEEE transactions on medical imaging 30.5 (2010): 1028-1041.
- Haldar, Justin P. "Low-rank modeling of local k-space neighborhoods (LORAKS) for constrained MRI." IEEE transactions on medical imaging 33.3 (2013): 668-681.
- Shin, Peter J., et al. "Calibrationless parallel imaging reconstruction based on structured low‐rank matrix completion." Magnetic resonance in medicine 72.4 (2014): 959-970.
- Jin, Kyong Hwan, Dongwook Lee, and Jong Chul Ye. "A general framework for compressed sensing and parallel MRI using annihilating filter based low-rank Hankel matrix." IEEE Transactions on Computational Imaging 2.4 (2016): 480-495.
- Wang, Shanshan, et al. "Accelerating magnetic resonance imaging via deep learning." 2016 IEEE 13th International Symposium on Biomedical Imaging (ISBI). IEEE, 2016.
- Hammernik, Kerstin, et al. "Learning a variational network for reconstruction of accelerated MRI data." Magnetic resonance in medicine 79.6 (2018): 3055-3071.
- Zhu, Bo, et al. "Image reconstruction by domain-transform manifold learning." Nature 555.7697 (2018): 487.
- Mardani, Morteza, et al. "Deep generative adversarial neural networks for compressive sensing MRI." IEEE transactions on medical imaging 38.1 (2018): 167-179.
- Aggarwal, Hemant K., Merry P. Mani, and Mathews Jacob. "Modl: Model-based deep learning architecture for inverse problems." IEEE transactions on medical imaging 38.2 (2018): 394-405.
- Yang, Guang, et al. "DAGAN: deep de-aliasing generative adversarial networks for fast compressed sensing MRI reconstruction." IEEE transactions on medical imaging 37.6 (2017): 1310-1321.
- Duan, Jinming, et al. "VS-Net: Variable splitting network for accelerated parallel MRI reconstruction." International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer, Cham, 2019.
- Han, Yoseob, Leonard Sunwoo, and Jong Chul Ye. "k-space deep learning for accelerated MRI." IEEE transactions on medical imaging (2019).
- Akçakaya, Mehmet, et al. "Scan‐specific robust artificial‐neural‐networks for k‐space interpolation (RAKI) reconstruction: Database‐free deep learning for fast imaging." Magnetic resonance in medicine 81.1 (2019): 439-453.
- Vasanawala, Shreyas S., et al. "Improved pediatric MR imaging with compressed sensing." Radiology 256.2 (2010): 607-616.
- Van Essen, D.C., et al: The wu-minn human connectome project: an overview. Neuroimage 80, 62-79 (2013).
- Schlemper, Jo, et al. "Nonuniform Variational Network: Deep Learning for Accelerated Nonuniform MR Image Reconstruction." International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer, Cham, 2019.
- Ronneberger, Olaf, Philipp Fischer, and Thomas Brox. "U-net: Convolutional networks for biomedical image segmentation." International Conference on Medical image computing and computer-assisted intervention. Springer, Cham, 2015.
- Schlemper, Jo, et al. "A deep cascade of convolutional neural networks for dynamic MR image reconstruction." IEEE transactions on Medical Imaging 37.2 (2017): 491-503.
- Wang, L-T., et al. "SSIM: a software levelized compiled-code simulator." Proceedings of the 24th ACM/IEEE Design Automation Conference. ACM, 1987.
- Wu B, Millane R, Watts R, Bones P. Applying compressed sensing in parallel MRI. In: Proceedings of the 16th Annual Meeting of ISMRM, vol. 1480 Toronto, Canada; 2008.
- Liu, Bo, Yi Ming Zou, and Leslie Ying. "SparseSENSE: application of compressed sensing in parallel MRI." 2008 International Conference on Information Technology and Applications in Biomedicine. IEEE, 2008.
- Coffey, Aaron M., Milton L. Truong, and Eduard Y. Chekmenev. "Low-field MRI can be more sensitive than high-field MRI." Journal of Magnetic Resonance 237 (2013): 169-174.
Figures
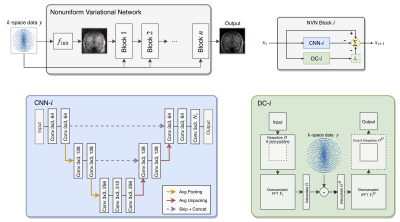
The architecture of Nonuniform Variational Network. Each block-$$$i$$$ has three components: CNN-$$$i$$$, DC-$$$i$$$ and $$$\lambda$$$. CNN-$$$i$$$ is a small network inspired by U-net to image de-aliasing, DC-$$$i$$$ is a non-Cartesian DC layer and $$$\lambda$$$ is the weighting term. The model has 5 blocks in total, which is end-to-end trainable. For more mathematically rigorous detail, refer to reference 28.
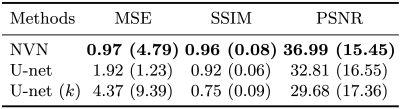
The quantitative results for the simulated data acquisition with system imperfections and the acceleration factor of 3.5. For each metric, mean and standard deviation are computed. The MSE's are scaled by 103. The proposed approach consistently outperformed the baseline approaches (U-net and k-space U-net). In particular, U-net alone was not powerful enough to correct all aliasing and $$$k$$$-space U-net struggled to converge, potentially due to the jittering of $$$k$$$-space samples.
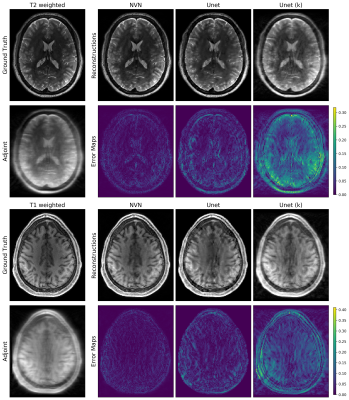
The reconstructions for the T1-weighted (bottom half) and T2-weighted images (top half) from the simulated data acquisition with system imperfections and the acceleration factor of 3.5. Top rows show the reconstructions and the bottom rows show the corresponding error maps. The reconstructions from NVN resulted in sharper images with highest data fidelities, owing to the data consistency blocks. U-net based methods could not resolve all aliasing artifacts correctly.
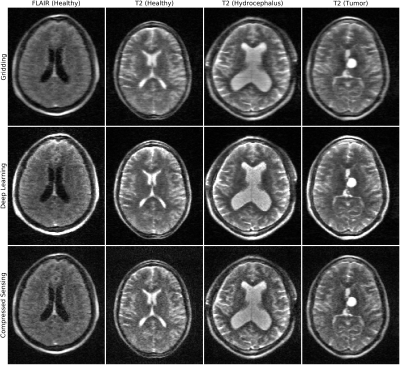
Linear (top), NVN (middle) and CS (bottom) reconstructions of prospectively undersampled low-field (64mT) images. From left: FLAIR and T2-weighted images from healthy controls, T2-weighted image from a patient with hydrocephalus, and T2-weighted image from a patient with a medial left thalamic cystic tumor and shunted hydrocephalus. Note the increased definition of NVN and CS, while NVN further reduced noise-like effect of CS.