4801
Segmentation of Intra-Tumour Distinct Metabolic Regions Using Chemical Exchange Saturation Transfer Imaging1Medical Biophysics, University of Toronto, Toronto, ON, Canada, 2Physical Sciences Platform, Sunnybrook Research Institute, Toronto, ON, Canada, 3Electrical and Computer Engineering, Lassonde School of Engineering, York University, Toronto, ON, Canada, 4Radiation Oncology, University of Toronto, Toronto, ON, Canada, 5Biological Sciences Platform, Sunnybrook Research Institute, Toronto, ON, Canada, 6Radiation Oncology, Sunnybrook Health Sciences Centre, Toronto, ON, Canada, 7Neurosurgery and Paediatric Neurosurgery, Medical University of Lublin, Lublin, Poland
Synopsis
Chemical exchange saturation transfer (CEST) is a promising MR contrast mechanism that has been shown to correlate with cancer metabolism and reveal regions of active tumour metabolism. However, the acquisition of CEST-weighted images is time consuming. In this study, computational methods including unsupervised learning were adapted to find the minimum number of CEST images required to segment the intra-tumour distinct metabolic regions accurately, and to find the number of different cell groups existing within a tumour. The results indicate that four intra-tumour regions can be segmented accurately using only CEST images acquired at 3.5 ppm and 2.0 ppm.
Introduction
Chemical exchange saturation transfer (CEST) is a promising MR contrast mechanism that has been shown to correlate with cancer metabolism and reveal regions of active tumour metabolism1. However, the acquisition of CEST-weighted images with radiofrequency saturation over a wide range of frequency offsets (“Z-spectra”) is time consuming. The goal of this study was to adapt computational methods including machine learning to find the minimum number of CEST images required to segment intra-tumour distinct metabolic regions accurately, and to find the number of different cell groups existing within a tumour using unsupervised clustering.Methods
Animal model: The model included 13 tumours grown in the hind limb of nude mice from two cell lines: DU145 human prostate adenocarcinoma (ATCC, Manassas, VA) and a radiation-resistant line generated by treatment of the former cell line with radiation mimicking a clinical treatment schedule1.
Dataset: Z-spectra for each tumour were acquired using magnetization transfer-prepared FLASH with steady-state saturation B1 of 0.5 and 2 µT. To perform clustering, for each saturation B1, images were used at the offset frequencies of two chemical groups: 3.5 ppm (amide) and 2.0 ppm (guanidinium)1.
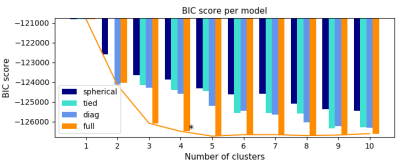
Data analysis: The optimal number of data points required for accurate segmentation of intra-tumour metabolic regions, as well as the optimal number of clusters was computed through minimizing the Bayesian information criterion (BIC) for clustering, and using the elbow method2 to select the number of clusters. The clustering algorithm was performed on independent components of the CEST images computed through independent component analysis (ICA). ICA is a linear transformation from the original feature space to a new feature space such that each of the new individual features are mutually independent3. The Gaussian mixture model (GMM) was used as the clustering method. GMM is a generative statistical method that applies probabilistic models to identify subpopulations within an overall population. The assumption behind GMM is that the data points within the d-dimensional population are coming from k different sources where each source is a Gaussian distribution4.
Results
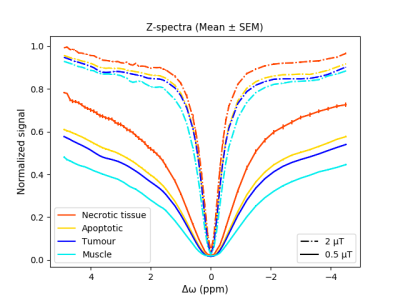
The optimization results indicate that the intra-tumour regions can be segmented using the independent components of two CEST images, where the optimal number of clusters is four (Fig. 1). As expected, the CEST data acquired at B1 = 0.5 µT is noisier, leading to over segmentation of the tumour. As such, clustering was performed on data acquired at B1 = 2 µT. Figure 2 depicts the Z-spectra averaged over all tumours for each cluster, and shows that the standard error of the mean (SEM) is small for all spectra, indicating small variance within each cluster. Qualitative comparison between the segmented image and the histology demonstrated that the intra-tumour regions can be segmented using only CEST images acquired at 3.5 ppm and 2.0 ppm (B1 = 2 µT). Figure 3 depicts an example of a tumour segmented with the proposed clustering approach.Discussion
The results indicate that, overall, the dataset includes four distinct metabolic regions. One is muscle (shown in cyan) while the rest are tumour sub-regions with different metabolic activities. Our hypothesis is that the least metabolically active cluster (red) corresponds to necrotic tumour, yellow is associated with apoptosis or partial necrosis while blue region corresponds to metabolically active parts of the tumour. The clustering performed on the CEST data (Fig. 3f) has good agreement with the histology (Fig. 3g).Conclusion
The results of this study suggest that CEST imaging and computational methods can potentially identify distinct metabolic regions within a tumour and characterize the intra-tumour heterogeneity. The proposed method can potentially be incorporated into cancer diagnosis and treatment planning workflows to develop more effective radiation dose maps.Acknowledgements
The authors would like to thank the CIHR (grant number PJT148660), and Terry Fox Research Institute (grant number 1083) for funding this project.References
1. Lam WW, Oakden W, Murray L, et al. Differentiation of normal and radioresistant prostate cancer xenografts using magnetization transfer-prepared MRI. Sci Rep. 2018. doi:10.1038/s41598-018-28731-0
2. Kodinariya TM, Makwana PR. Review on determining number of Cluster in K-Means Clustering. IJARCSMS. 2013.
3. Hyvärinen A, Oja E. Independent component analysis: Algorithms and applications. Neural Networks. 2000. doi:10.1016/S0893-6080(00)00026-5
4. Reynolds D. Gaussian Mixture Models. In: Encyclopedia of Biometrics. ; 2015. doi:10.1007/978-1-4899-7488-4_196
Figures