4069
Fetal Brain-Heart-Placental Interactions with Acute Hypoxia Challenge in Genetic Mouse Models of Hypoplastic Left Heart Syndrome with in utero 4D Dynamic MRI1Biomedical Imaging Research Institute, Cedars-Sinai Medical Center, Los Angeles, CA, United States, 2Developmental Biology, University of Pittsburgh, Pittsburgh, PA, United States, 3Rangos Research Center Animal Imaging Core, Children's Hospital of Pittsburgh of UPMC, Pittsburgh, PA, United States
Synopsis
The objective of this study is to develop 4D time-resolved high-resolution in utero MRI for simultaneous interrogation of placenta, heart and brain development and function in genetic mouse models of hypoplastic left heart syndrome. The structural and functional imaging of the fetal heart, brain, and placenta can be correlated with genotype. In addition, temporal and spatial characteristics of BOLD responses to acute hypoxia challenge can be derived.
Introduction
Congenital heart defects (CHD) are the most common birth defects, affecting nearly 1% of live births. The survival rates for CHD patients have greatly improved with surgical advances, however, CHD patients suffer higher incidences of neurocognitive impairment. Intrauterine hypoxia is thought to be the key determinant of poor neurocognitive development in CHD. However, preliminary clinical trials with maternal hyperoxygenation to improve fetal neurodevelopment fail to deliver the favorable outcome. On the other hand, placental structural and vascular abnormalities have been associated with hypoplastic left heart syndrome (HLHS) and other CHD, indicating the strong placental-fetal heart interactions. Furthermore, the placental weight was found to be strongly correlated with the cerebral growth in CHD fetuses, indicating the intimate placental-fetal brain relationship. A critical unanswered question is whether the poor neurodevelopmental outcome in CHD is driven by abnormal placenta, hemodynamic perturbations arising from the structural heart defects, cumulative injury from hypoxia and surgery, or primarily driven by patient intrinsic/genetic factors. These questions cannot be answered in patients because surgical intervention is required for patient survival. Mouse models harboring genetic mutations associated with CHD are invaluable model systems for investigating mechanistic insight into the structure-function relationship between placenta, fetal heart, and fetal brain development in utero. The goal of this study is to develop 4D dynamic MRI in utero for simultaneous interrogation of placental, cardiovascular and neurodevelopment in genetic mouse models of HLHS.Methods
A. 4D Dynamic Fetal MRI:
We use a hybrid1 low-rank2 and sparse3 model to perform 4D BOLD imaging in utero. The low-rank model expresses the image $$$\rho(\mathbf{r},t)$$$ (for voxel location $$$\mathbf{r}$$$ and time $$$t$$$) as:$$\rho(\mathbf{r},t)=\sum_{\ell=1}^L\psi_\ell(\mathbf{r})\varphi_\ell(t),$$where $$$\{\psi_\ell(\mathbf{r})\}_{\ell=1}^L$$$ are $$$L$$$ spatial coefficient maps and where $$$\{\varphi_\ell(t)\}_{\ell=1}^L$$$ are $$$L$$$ temporal basis functions. In matrix form, this becomes $$$\mathbf{X=\Psi\Phi}$$$, where $$$X_{ij}=\rho(\mathbf{r}_i,t_j)$$$, $$$\mathit{\Psi}_{ij}=\psi_j(\mathbf{r}_i)$$$, and $$$\mathit{\Phi}_{ij}=\varphi_i(t_j)$$$. We additionally model $$$\rho(\mathbf{r},t)$$$ as being transform sparse, specifically in the spatial-spectral domain (i.e., we model $$$\mathcal{F}_t\{\rho(\mathbf{r},t)\}$$$ as sparse, where $$$\mathcal{F}_t$$$ is the temporal Fourier transform). This approach exploits both image correlation and transform sparsity to allow high spatiotemporal resolution imaging. We perform image reconstruction in two steps: first determining the temporal basis $$$\mathbf{\Phi}$$$ from the singular value decomposition (SVD) of interleaved training data, and second determining the spatial coefficient maps $$$\mathbf{\Psi}$$$ by fitting the temporal basis to the remainder of the imaging data. This second step is calculated according to$$\mathbf{\Psi}=\arg\min_\mathbf{\Psi}\|\mathbf{d}-E(\mathbf{\Psi\Phi})\|_2^2+\lambda\|\mathbf{\Psi\Phi{F}}\|_1$$where $$$\mathbf{d}$$$ are the measured data, $$$E$$$ is the encoding operator comprising spatial encoding and undersampling, and $$$\mathbf{F}$$$ is the temporal Fourier transform.
B. Animal Model:
Digenic mouse model of HLHS4 with both Pcdha9 and Sap130 gene mutations were used in this study. The pcdha9 gene encodes protocadherinA9 mediating cell-cell adhesion, whereas Sap130 encodes Sin3A-associated protein 130, a member of the histone deacetylase (HDAC) complex mediating chromatin repression. Mouse fetuses with homozygous mutations in both pcdha9 and Sap130 exhibit HLHS. Some HLHS mutant mice show microcephaly and holoprosencephaly, similar to brain defects seen in HLHS patients. Mice heterozygous for both pcdha9 and Sap130 do not display HLHS phenotype. In this study, double heterozygous adult breeding pairs were used to generate fetuses with various genotypes within the same pregnancy.
Results
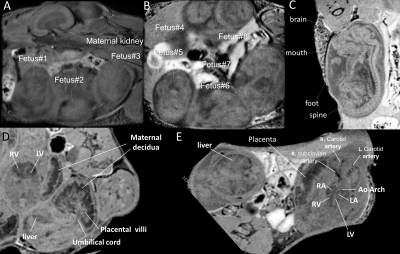
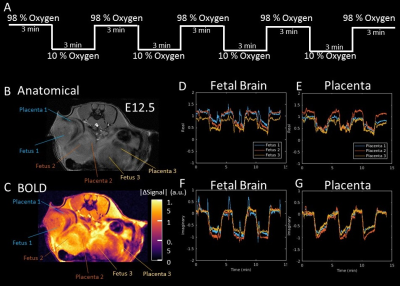
The two key features of this fetal MRI approach are (1) the ability to express incoherent fetal and maternal motion with reduced degrees of freedom; and (2) a sparse sampling scheme to accelerate acquisition and increase temporal resolution. This allows assessments of anatomical structures and the hemodynamic function and its relationship with the developing fetal brain. Fig1 shows single imaging planes at one time point of the 4D MRI performed in utero of a pregnant female mouse on embryonic day E15.5. Multiple fetuses can be imaged without motion artifact. Fetal brain, heart, liver, and placental structure can be identified. Oscillating acute hypoxia challenge (Fig.2A) every 3 minutes was subjected to a double heterozygous female mouse carrying fetuses on E12.5. 3 out of the 8 embryos can be seen on the imaging plane shown (Fig. 2B,C). The degrees of the blood oxygenation level dependent (BOLD) signal changes responding to the oscillating hypoxia changes are color-coded for the 3 fetuses (Fig.2C). The temporal BOLD responses for the fetal brains (Fig. 2 D, F) and placenta (Fig.2 E,G), expressed as real (Fig. 2 DE) and imaginary (Fig.2FG) components. These 3 fetuses with different genotypes displayed different degrees of BOLD responses and temporal dynamic characteristics which are able to be captured by the 4D MRI.Conclusion
Our 4D time-resolved MRI can capture the dynamic BOLD signals with high spatial and temporal resolution for fetal brain and placenta to correlate with different genotypes.Acknowledgements
The authors thank Nathan Salamacha, Cassandra Slover, Samuel Wyman, Lauren Myers, and Cullen Yang, for assisting with managing animals.References
1 Zhao B et al. IEEE-TMI 2012.
2 Liang Z-P. IEEE-ISBI 2007.
3 Lustig M et al. MRM 2007.
4 Liu, et al. Nature Gen. 2017.
Figures