3532
Novel HARDI-ZOOMit protocol detects changes in spinal cord microstructure in patients with asymptomatic non-myelopathic degenerative cervical spinal cord compression1Department of Neurology, Palacký University, Olomouc, Czech Republic, 2Department of Neurology, University Hospital Olomouc, Olomouc, Czech Republic, 3Department of Pediatrics, University of Minnesota, Minneapolis, MN, United States, 4Department of Biomedical Engineering, University Hospital Olomouc, Olomouc, Czech Republic, 5Central European Institute of Technology, Masaryk University, Brno, Czech Republic, 6Department of Neurology, University Hospital Brno, Brno, Czech Republic, 7Department of Medicine III, Clinical Division of Endocrinology and Metabolism, Medical University of Vienna, Vienna, Austria, 8High Field MR Centre, Medical University of Vienna, Vienna, Austria, 9Center for Magnetic Resonance Research, University of Minnesota, Minneapolis, MN, United States, 10Institute of Biomedical Engineering, Polytechnique Montreal, Montreal, QC, Canada
Synopsis
Detection of degenerative cervical spinal cord compression (DCSCC) at early stage that causes spinal cord (SC) microstructure disruption is limited when current imaging techniques are used. Early detection of damage to SC can predict symptomatic degenerative cervical myelopathy (DCM). We are presenting novel HARDI-ZOOMit protocol (High Angular Resolution Diffusion Imaging + syngo ZOOMit sequence), which is able to detect changes in microstructural diffusion MRI (dMRI) parameters (e.g. fractional anisotropy – FA, mean diffusivity – MD) in asymptomatic DCSCC. Results obtained with HARDI-ZOOMit protocol reveal higher sensitivity when compared to clinical RESOLVE (REadout Segmentation Of Long Variable Echo trains) protocol.
Introduction
Degenerative cervical spinal cord compression (DCSCC) may result in changes of microstructural diffusion MRI (dMRI) parameters in patients with symptomatic DCM in comparison to DCSCC patients without clinical signs and myelopathy symptoms (i.e. non-myelopathic degenerative cervical spinal cord compression – NMDCSCC)1. Quantitative MRI including dMRI detected microstructural SC lesion in NMDCSCC2. We have introduced a novel HARDI-ZOOMit (High Angular Resolution Diffusion Imaging) protocol visualizing crossing fibers from in-vivo human cervical data3 and are employing it to detect differences between NMDCSCC patients (with DSCCs mostly in C4-C6 segments) and healthy controls.
Methods
High-resolution sagittal T2-weighted and axial T2*-weighted anatomical scans followed by two independent axial dMRI acquisitions (i.e. HARDI-ZOOMit and RESOLVE protocols) were acquired on 3T Siemens Prisma MR scanner using 64-channel head/neck and 32-channel spine coils. Five young healthy volunteers (3 males, 27.6±1.8 y.o., measured twice), 15 age-matched healthy controls (5 males, 50.1±9.6 y.o.) and 20 NMDCSCC patients (11 males, 58.9±9.5 y.o.) were enrolled in the study approved by the local ethical committee.
T2-weighted sagittal images were acquired to cover the whole cervical SC with 30 contiguous slices using turbo spin-echo sequence with TR=8640ms (repetition time), TE=98ms (echo time), 4 averages, GRAPPA=2, voxel size 0.28×0.28×1.3mm3 (original 0.56×0.56×1.3mm3).
T2*-weighted axial images were acquired to cover the C3-C7 levels with 42 contiguous slices using Multi-Echo Data Image Combination sequence with TR=778ms, TE=17ms (4 echoes), 2 averages, voxel size 0.35×0.35×2.5mm3 (original 0.7×0.7×2.5mm3).
HARDI-ZOOMit dMRI covered the C3-C7 levels with 35 contiguous axial slices with TR=6700ms, TE=73ms, original voxel size 1.3×1.3×3mm3 re-sampled at 0.65×0.65×3mm3 in the Fourier domain. Sixty-three diffusion-weighted images (42 gradient directions with b=1000smm-2 and 21 directions with b=550smm-2) and 7 images (b0) with b=0smm-2 were collected with anterior-posterior (AP) phase encoding. Five additional b0 images were acquired using PA encoding, acquisition time TA=12min46s. For the sub-sets of above mentioned participants, the protocol was repeated with turned off interpolation.
RESOLVE dMRI consisted of two identical measurements with opposite AP/PA encodings. For both, 30 diffusion weighted images with b=650smm-2 and 5 b0 images were collected covering C3-C7 levels with 30 contiguous axial slices with TR=4500ms, TE1=50ms, TE2=77ms, voxel size 1.1×1.1×3.3mm3, TA=16min16s.
The acquired data were processed with Spinal Cord Toolbox 3.2.34, Advanced Normalization Tools 2.1.05 and FSL 5.0.106 software libraries wrapped within in-house made shell and MATLAB (MathWorks, USA) scripts. Susceptibility, eddy current and motion artifacts7,8 were minimized from dMRI data and diffusion tensor imaging (DTI) model9 was fitted. T2*-w axial scan was co-localized based on T2-w sagittal scan. Final SC and white matter (WM)/gray matter (GM) segmentations10,11 were performed in T2*-w axial space with simultaneous segment labeling12. DTI derived metrics (i.e. fractional anisotropy FA and mean diffusivity MD) of all protocols were warped into T2*-w axial space, and single-subject C3-C6 WM/GM parameters (i.e. mean, median, mode, standard deviation, skewness and kurtosis of FA/MD distribution) were estimated. Wilcoxon signed-rank test evaluated differences between single-subject parameters of different groups. Pairwise differences with p<0.01667 (≈pFWE<0.05) were considered significant.
Results
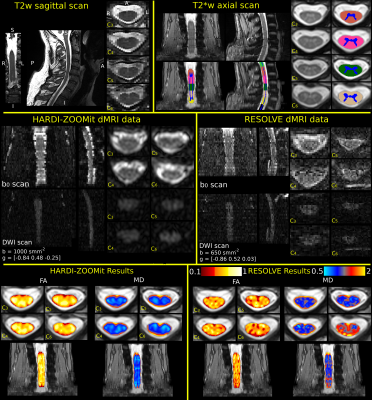
The proposed HARDI-ZOOMit protocol is about 3.5 minutes faster than
RESOLVE protocol. Example of single-subject acquired data with
visualization of segmentation and FA with MD results
is shown in Fig.
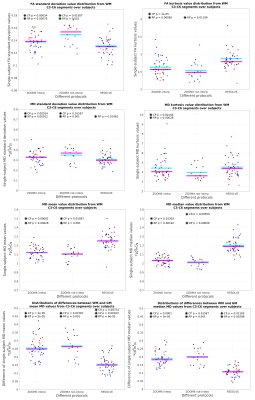
1. In WM, significant differences between age-comparable control
and patient groups were observed for standard deviation of FA and MD,
kurtosis of FA and MD, mean/median of MD for our novel HARDI-ZOOMit protocol (Fig. 2). RESOLVE protocol did not detect those differences (Fig.
2). All protocols demonstrate significant differences for WM-GM
subtraction of MD means/medians over all three groups. HARDI-ZOOMit protocol still has the lowest p-values (Fig. 2).
FA WM kurtosis and WM-GM
mean subtraction
demonstrate the lowest control-patient p=2e-5.
All
other supra-threshold p-values are listed in Fig.
2.
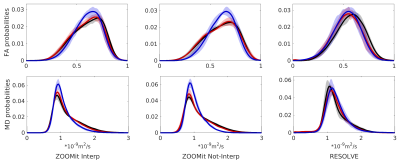
Differences between WM FA/MD parameters over groups are
observable within smoothed group-averaged probability density
functions for HARDI-ZOOMit protocol (Fig. 3). No significant
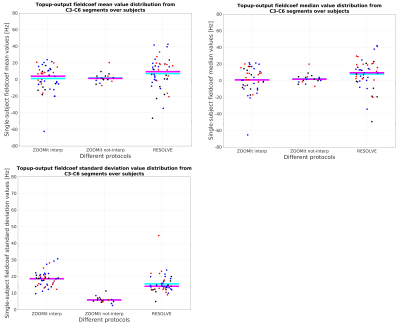
changes for C3-C6 GM were observed. Susceptibility artifacts seem to
be in the same orders for both protocols with lower frequency
deviations for HARDI-ZOOMit (Fig. 4).
Discussion
Significant differences in C3-C6 WM may be interpreted as microstructural changes over imaged tract sections; future studies will investigate more spatially focused tract-specific changes using dMRI techniques. Results of non-interpolated HARDI-ZOOMit have similar distribution as the interpolated version (Fig. 2). Lack of significant GM changes may reflect the segmental nature of GM damage, which can be obscured by averaging over the whole C3-C6 region.Conclusion
We present a novel dMRI protocol capable of detecting changes in DTI derived parameters for patients at early asymptomatic stage of DCSCC. The protocol has a potential to predict development of DCM at early microstructural presymptomatic stage.Acknowledgements
We acknowledge the core facility Multimodal and Functional Imaging Laboratory, Masaryk University, CEITEC supported by the MEYS CR (LM2015062 Czech-BioImaging). This research was supported and funded by the Czech Health Research Council grant n. NV18-04-00159, and by the Ministry of Health of the Czech Republic project for conceptual development in research organizations, ref. 65269705 (University Hospital, Brno, Czech Republic). CL is partly supported by NIH grants P41 EB015894 and P30 NS076408. AS has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No 794986.References
1. Keřkovský M, Bednařík J, Dušek L, et al. Magnetic Resonance Diffusion Tensor Imaging in Patients With Cervical Spondylotic Spinal Cord Compression. Spine. 2012;37:48–56.
2. Martin A R, De Leener B, Cohen-Adad J, et al. Can microstructural MRI detect subclinical tissue injury in subjects with asymptomatic cervical spinal cord compression ? A prospective cohort study. BMJ Open. 2018;8:e019809.
3. Labounek R, Valošek J, Horák T, et al. Fast in vivo high-resolution diffusion MRI of the human cervical spinal cord microstructure. in IFMBE Proceedings: World Congress on Medical Physics & Biomedical Engineering. 2019;68(1):3–7.
4. De Leener B, Lévy S, Dupont S M, et al. SCT: Spinal Cord Toolbox, an open-source software for processing spinal cord MRI data. Neuroimage. 2017;145:24–43.
5. Avants B B, Tustison N & Song G. Advanced Normalization Tools (ANTs). Insight J. 2009;2:1–35.
6. Jenkinson M, Beckmann C F, Behrens T E J, Woolrich M W & Smith S M. Fsl. Neuroimage. 2012;62:782–790.
7. Andersson J L R, Skare S & Ashburner J. How to correct susceptibility distortions in spin-echo echo-planar images: Application to diffusion tensor imaging. Neuroimage. 2003;20:870–888.
8. Andersson, J. L. R. & Sotiropoulos, S. N. An integrated approach to correction for off-resonance effects and subject movement in diffusion MR imaging. Neuroimage 125, 1063–1078 (2016).
9. Basser P J, Mattiello J & LeBihan D. MR diffusion tensor spectroscopy and imaging. Biophys. J. 1994;66:259–67.
10. Gros C, De Leener B, Badji A, et al. Automatic segmentation of the spinal cord and intramedullary multiple sclerosis lesions with convolutional neural networks. Neuroimage. 2018;184:901–915.
11. Perone C S, Calabrese E & Cohen-Adad J. Spinal cord gray matter segmentation using deep dilated convolutions. Sci. Rep. 2018;8:1–13.
12. De Leener B, Cohen-Adad J & Kadoury S. Automatic Segmentation of the Spinal Cord and Spinal Canal Coupled with Vertebral Labeling. IEEE Trans. Med. Imaging 2015;34:1705–1718.
Figures