3409
HARDI denoising with mean-curvature enhancement PDE on SE(3)1SCIL, Université de Sherbrooke, Sherbrooke, QC, Canada, 2CASA, Department of Mathematics and Computer Science, Eindhoven University of Technology, Eindhoven, Netherlands
Synopsis
We present a new High Angular Resolution Diffusion-weighted Imaging (HARDI) mean-curvature enhancement (MCE) on the homogeneous space of positions and orientations, embedded in the rigid body motion group SE(3). Its potential for crossing-preserving enhancement of fiber orientation distribution (FOD) fields is demonstrated. Compared to previous partial differential equation (PDE) enhancements on SE(3), denoising FOD fields with MCE better preserve boundaries, resulting in a lower overall angular error.
Introduction
Anisotropic filtering was first introduced to reduce noise from images without blurring sharp edges1,2. These edge-preserving smoothing PDE encourage the conductivity along contours, increasing spatial consistency, while still preserving angular features (boundaries, corners, contour curves). As an alternative to these methods, curvature based filtering was proposed to reduce noise in images and discrete surfaces3,4,5.
For diffusion-weighted MRI (DW-MRI), numerous filtering methods have been proposed to increase the local coherence of Diffusion Tensor Image (DTI)6,7,8. Since, DTI is unable fully represent the diffusion signal in crossing regions, known to occur in most of the white matter [9,10], advanced methods were proposed to increase spatial continuity of multi-tensor fields11,12,13 or FOD fields14,15. Multiple approaches for HARDI denoising have also been compared by Daducci et al16.
Previous works exposed the advantage of PDE on SE(3) for crossing-preserving filtering17. This representation enables the processing of spatial and angular components together, since the underlying metric combines both, i.e. high diffusivity in a certain orientation should persist in the spatial domain along this orientation. Elaborations of these concepts on $$$\mathbb{R}^3 \rtimes \mathbb{S}^2$$$ led to contextual enhancement (CE) and Perona-Malik enhancement (PME) for 2D or 3D orientation score images and DW-MRI18,19.
Method
In this work, we adapted an anisotropic filtering approach for HARDI, to enhance the alignment of elongated structures while maintaining crossing angles. This enhancement follows previous research on mean-curvature ($$$\kappa$$$) enhancement on 2D images ($$$I(\mathbf{x}), \, \mathbf{x} \in \mathbb{R}^2$$$) or manifold2,3,4,5:$$\frac{\partial J(\mathbf{x},t)}{\partial t}= \|\nabla J(\mathbf{x},t)\| \kappa(\mathbf{x},t)= \|\nabla J(\mathbf{x},t)\| \text{div} \left(\frac{\nabla J(\mathbf{x},t)}{\|\nabla J(\mathbf{x},t)\|} \right)\\
J(\mathbf{x}, 0)= I(\mathbf{x}), \quad t>0$$
For HARDI, the image domain ($$$\mathbb{R}^3$$$) was extended with a sphere ($$$\mathbb{S}^2$$$), considering that FODs generate a spatial field of spherical function ($$$U(\mathbf{x},\mathbf{n}),\, (\mathbf{x},\mathbf{n}) \in \mathbb{R}^3 \times \mathbb{S}^2$$$). Hence, the MCE equation was adapted to the Lie group SE(3) (or rather Lie group quotient $$$\mathbb{R}^3 \rtimes \mathbb{S}^2$$$) with the use of Left-invariant derivative19,20,21:
$$\frac{\partial W(\mathbf{x},\mathbf{n},t)}{\partial t}= \|\nabla_{sr} W(\mathbf{x},\mathbf{n}, t) \| \left(D_{\mathbf{x}} (\mathbf{n} \cdot \nabla_{\mathbb{R}^3})\left(\frac{(\mathbf{n} \cdot \nabla_{\mathbb{R}^3}) W(\mathbf{x},\mathbf{n},t)}{\|\nabla_{sr} W(\mathbf{x},\mathbf{n},t)\|}\right) + D_{\mathbf{n}}\, \textrm{div}_{\mathbb{S}^2}\left(\frac{\nabla_{\mathbb{S}^2} W(\mathbf{x},\mathbf{n},t)}{\|\nabla_{sr} W(\mathbf{x},\mathbf{n},t)\|}\right) \right)\\
\|\nabla_{sr} W(\mathbf{x},\mathbf{n}, t) \| = \sqrt{\frac{D_{\mathbf{x}}}{D_{\mathbf{n}}}\|(\mathbf{n} \cdot \nabla_{\mathbb{R}^3}) W(\mathbf{x},\mathbf{n},t)\|^2 + \|\nabla_{\mathbb{S}^2} W(\mathbf{x},\mathbf{n},t)\|^2}\\
W(\mathbf{x},\mathbf{n}, 0)= U(\mathbf{x},\mathbf{n}), \quad t>0$$
Evaluation
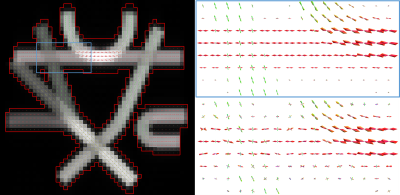
To evaluate various SE(3) filtering techniques, we used the ISBI-HARDI challenge synthetic phantom22 at SNR10 and computed the FOD field with constrained spherical deconvolution (CSD)23 (Figure #1). To analyze results, the noisy FOD field was filtered with different parameters on 40 iterations ($$$t=4, dt=0.1$$$): angular conductivity $$$D_{\mathbf{n}} = 0.001, 0.002, 0.004$$$ and PME sensitivity $$$K = 0.1, 0.2, 0.4, 0.8$$$. A constant rescaling of both $$$D_{\mathbf{x}}$$$ and $$$D_{\mathbf{n}}$$$ correspond to a scaling of the diffusion time ($$$t, dt$$$), therefore we kept the spatial conductivity $$$D_{\mathbf{x}} = 1$$$ fixed.
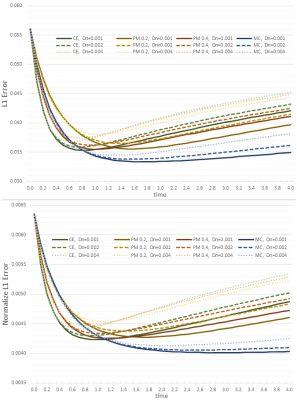
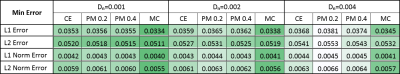
Quantitatively, the L1 and L2 absolute error, defined as the distance to the noiseless FOD, was computed at each timestep, for all SE(3) enhancements. A second version of both L1 and L2 error was included to assess only the orientational error, where at each position the FOD was normalized.
Qualitatively, a standard in-vivo HARDI acquisition (49 directions at b-value 1000) was included to visually compare results.
Result
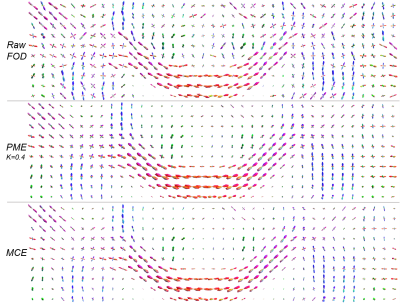
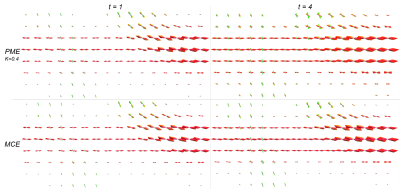
Figure #2 illustrates filtering results over the synthetic phantom, comparing the proposed CME to the PME. Since there was no perceptual difference between the CE and PME, only one of them is displayed. The L1 error progression for all methods are shown in Figures #3-4, the L2 error followed the same trend. The resulting enhanced FOD, over a standard in-vivo dataset, can be observed in Figure #5. Only PM results with sensitivity $$$K = 0.2, 0.4$$$ are presented as they showed best performance compared to $$$K = 0.1, 0.8$$$.Discussion
As illustrated in Figure #2, all methods are able to remove spurious peaks, and preserve crossings. Contextual enhancement and Perona-Malik methods reduce noise from FOD field within few iterations, but afterward tend to blur edges ($$$t > 2$$$), causing the error to increase (see Figures #3-4). The proposed MCE is able to further decrease the error before over-smoothing. In addition, the reduced normalized error, combined with the smaller slope, (Figure #4) makes the proposed filtering very appealing when the FOD field is only used as an angular probability, e.g. for tractography. In Figure #5, sharper and more consistent FOD is obtained with MCE.Conclusion
Mean-curvature enhancement, with SE(3) framework, can denoise HARDI both spatially and angularly. The proposed method is particularly good to recover the angular distribution of FODs. Parameters and diffusion time can be optimized to maximize FOD denoising while preserving sharp edges. In future work, it would be interesting to adapt the MCE for contextual interpolation/upsampling or to regularize the CSD process.Acknowledgements
We would like to thank Dr. A. Roebroeck from the Faculty of Psychology & Neuroscience, Maastricht University, for providing us with the in-vivo data. Supported by grants from Université de Sherbrooke Research Chair, NSERC Discovery Grant, European Research Council under the European Community’s Seventh Framework Programme (FP7/2007-2013), ERC grant Lie Analysis (agr. nr. 335555).References
1. P. Perona and J. Malik. Scale space and edge detection using anisotropic diffusion.IEEE Trans. Patt. Anal. Mach. Intell., 12:629–639, 1990.
2. El-Fallah, Adel I., and Gary E. Ford. "The evolution of mean curvature in image filtering." Image Processing, 1994. Proceedings. ICIP-94., IEEE International Conference. Vol. 1. IEEE, 1994.
3. Desbrun, Mathieu, et al. "Implicit fairing of irregular meshes using diffusion and curvature flow." Proceedings of the 26th annual conference on Computer graphics and interactive techniques. ACM Press/Addison-Wesley Publishing Co., 1999.
4. Citti, Giovanna, et al. "Sub-Riemannian mean curvature flow for image processing." SIAM Journal on Imaging Sciences 9.1 (2016): 212-237.
5. Gong, Yuanhao, and Ivo F. Sbalzarini. "Curvature filters efficiently reduce certain variational energies." IEEE Transactions on Image Processing 26.4 (2017): 1786-1798.
6. Tschumperlé, David, and Rachid Deriche. "Diffusion tensor regularization with constraints preservation." Computer Vision and Pattern Recognition, 2001. CVPR 2001. Proceedings of the 2001 IEEE Computer Society Conference on. Vol. 1. IEEE, 2001.
7. Hamarneh, Ghassan, and Judith Hradsky. "Bilateral filtering of diffusion tensor magnetic resonance images." IEEE Transactions on Image Processing 16.10 (2007): 2463-2475.
8. Baust, Maximilian, et al. "Combined Tensor Fitting and TV Regularization in Diffusion Tensor Imaging based on a Riemannian Manifold Approach." (2016).
9. Vos, Sjoerd B., et al. "The influence of complex white matter architecture on the mean diffusivity in diffusion tensor MRI of the human brain." Neuroimage 59.3 (2012): 2208-2216.
10. Jeurissen, Ben, et al. "Investigating the prevalence of complex fiber configurations in white matter tissue with diffusion magnetic resonance imaging." Human brain mapping 34.11 (2013): 2747-2766.
11. Sotiropoulos, Stamatios N., et al. "A regularized two-tensor model fit to low angular resolution diffusion images using basis directions." Journal of Magnetic Resonance Imaging 28.1 (2008): 199-209.
12. Taquet, Maxime, Benoit Scherrer, and Simon K. Warfield. "A Framework for the Analysis of Diffusion Compartment Imaging (DCI)." Visualization and Processing of Higher Order Descriptors for Multi-Valued Data. Springer International Publishing, 2015. 271-297.
13. Cabeen, Ryan P., and David H. Laidlaw. "Bilateral filtering of multiple fiber orientations in diffusion MRI." Computational Diffusion MRI. Springer International Publishing, 2014. 193-202.
14. Reisert, Marco, and Henrik Skibbe. "Fiber continuity based spherical deconvolution in spherical harmonic domain." International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer, Berlin, Heidelberg, 2013.
15. Canales-Rodríguez, Erick J., et al. "Spherical deconvolution of multichannel diffusion MRI data with non-Gaussian noise models and spatial regularization." PloS one 10.10 (2015): e0138910.
16. Daducci, Alessandro, et al. "Quantitative comparison of reconstruction methods for intra-voxel fiber recovery from diffusion MRI." IEEE transactions on medical imaging 33.EPFL-ARTICLE-183667 (2014): 384-399.
17. Duits, Remco, and Erik Franken. "Left-invariant diffusions on the space of positions and orientations and their application to crossing-preserving smoothing of HARDI images." International Journal of Computer Vision 92.3 (2011): 231-264.
18. Creusen, Eric, et al. "Numerical schemes for linear and non-linear enhancement of DW-MRI." Numerical Mathematics: Theory, Methods and Applications 6.1 (2013): 138-168.
19. Portegies, Jorg M., et al. "Improving fiber alignment in HARDI by combining contextual PDE flow with constrained spherical deconvolution." PloS one 10.10 (2015): e0138122.
20. Portegies, J. M., and R. Duits. "New exact and numerical solutions of the (convection-) diffusion kernels on SE (3)." arXiv preprint arXiv:1604.03843 (2016).
21. Janssen, Michiel HJ, et al. "The Hessian of axially symmetric functions on SE (3) and application in 3D image analysis." International Conference on Scale Space and Variational Methods in Computer Vision. Springer, Cham, 2017.
22. Neher, Peter F., et al. "Fiberfox: facilitating the creation of realistic white matter software phantoms." Magnetic resonance in medicine 72.5 (2014): 1460-1470.
23. Tournier, J-Donald, Fernando Calamante, and Alan Connelly. "Robust determination of the fibre orientation distribution in diffusion MRI: non-negativity constrained super-resolved spherical deconvolution." Neuroimage 35.4 (2007): 1459-1472.
Figures