3073
MRI-based deep learning prediction of high amino acid uptake region to improve survival prediction in patents with glioblastoma: A 3D U-net study with deeply learned inter-scanner multi-modal MRI and alpha-[11C]-methyl-L-tryptophan (AMT) PETJeong-Won Jeong1,2, Min-Hee Lee2,3, Flora John2,3, Sandeep Mittal4,5, and Csaba Juhasz1,2
1Pediatrics, Neurology, Translational Neuroscience Program, Wayne State University, Detroit, MI, United States, 2Translational Imaging Laboratory, PET center, Children's Hospital of Michigan, Detroit, MI, United States, 3Pediatrics and Neurology, Wayne State University, Detroit, MI, United States, 4Neurosurgery and Oncology, Wayne State University, Detroit, MI, United States, 5Karmanos Cancer Institute, Detroit, MI, United States
Synopsis
Previous studies found that high amino acid uptake measured by alpha-[11C]-methyl-L-tryptophan (AMT)-PET can accurately detect glioblastoma cell infiltration both in enhancing and non-enhancing tumor portions. However, AMT-PET is not widely available for clinical use. This study explores a novel U-Net which can accurately detect high tryptophan uptake glioblastoma regions using clinical multi-modal MRI data. The resulting U-Net led to 0.85±0.08 sensitivity and 0.99±0.00 specificity to predict AMT-PET tumor regions showing significant negative correlation with survival period, suggesting that an end-to-end deep learning of multi-modal MRI data may be effective for survival prediction of glioblastoma patient without the need of AMT-PET.
Introduction
Glioblastomas are the most deadly primary brain tumors, and their initial treatment (surgery followed by radiation), based on clinical MRI, can miss tumor portions infiltrating to adjacent brain regions. Previous studies1,2 found that high amino acid uptake measured by alpha-[11C]-methyl-L-tryptophan (AMT)-PET can accurately detect glioblastoma cell infiltration both in enhancing and non-enhancing tumor portions. Thus, AMT uptake is a strong and independent imaging marker of the metabolically active tumor region which can be powerful for survival prediction3,4. However, amino acid (including AMT) PET is not widely available for clinical use. This study explores a novel end-to-end deep learning framework which can accurately detect high tryptophan uptake glioblastoma regions (the metabolic tumor volume) using clinical multi-modal MRI data obtained from two different imaging protocols.Methods
Preoperative contrast-enhanced T1, non-contrast T2/FLAIR, DWI apparent diffusion coefficient, and AMT-PET images were analyzed in 21 patients with newly diagnosed glioblastoma (mean age: 58 years) where 12 and 9 patients were scanned by different multi-modal MRI protocols summarized in Table 1. All patients underwent the same AMT-PET scanning protocol1-4. Multi-modal MRI images were intensity-scaled by global mean, spatially co-registered and resampled at the same resolution (1mm×1mm×1mm). A binary mask of the metabolically active tumor was obtained as the ground truth from AMT-PET by applying a previously established threshold of 1.65 tumor/normal cortex ratio3,4. To investigate the effect of different MRI protocols on overall performance of the proposed deep learning framework, three different 3D U-net systems5,6 (1st: Siemens, 2nd: Philips, and 3rd: Siemens and Philips) were separately implemented using Google TensorFlow library (www.tensorflow.org) with 4 layers of the encoding and decoding paths. Each U-net system was designated to deeply learn nonlinear voxel-wise relationship between “given input: multi-modal MRI” and “targeted output: AMT-PET tumor mask” where dice similarity coefficient (DSC) was used as a measure of detectability and optimized by back-propagating a loss function through the U-net. To test if the AMT PET-learned MRI-based tumor volume (i.e., output of multi-modal MRI U-Net) outperforms clinically used contrast-enhancement for survival prediction, overall survival (days; reliable data available in 19 patients) was correlated with: 1) Contrast-enhancing tumor volume (mm3) from the T1-Gad image and 2) AMT PET-learned MRI-based tumor volume.Results
Data augmentation was performed to generate 2100 study samples (i.e., 100 augmentations per patient) by applying random affine transformation to the original data of 21 patients (70%/30% for training/validation set). After 5000 iterations, DSC values of three U-Net systems: Siemens/Philips/Siemens+Philips reached 0.98(0.98)/0.99(0.99)/0.98(0.98) in the training (validation) set, respectively (Table 2). At the voxel level, the resulting U-Net models led to 0.87±0.05/0.92±0.05/0.85±0.08 sensitivity, 0.99±0.0/0.99±0.00/0.99±0.00 specificity, 0.86±0.06/0.84±0.04/0.81±0.08 positive predictive value and 0.99±0.00/0.99±0.00/0.99±0.00 negative predictive value (Table 2). Figure 1 presents a representative example to predict glioblastoma volume using the proposed U-Net system with multi-modal MRI data. It is clear the proposed U-Net system could accurately identify two structural lesions (U-Net determined glioma, blue contours) which were spatially well-matched with ground truth (AMT determined glioma, red contours). The AMT-PET-learned MRI-based tumor volume was significantly larger than the contrast-enhancing volume (mean: 41.5×103 mm3 vs. 18.9×103 mm3, p=0.001 in paired t-test). Both volume types showed a trend for a negative correlation with overall survival, with the relationship showing a non-linear component (Fig. 2): an exponential fit showed a significant negative relationship with the AMT-PET-learned MRI tumor volume (R2=0.24, p=0.03) but was not significant with the contrast-enhancing volume (R2=0.14, p=0.14). The linear correlation showed a trend for both volumes (p=0.11 and 0.10, respectively).Discussion
This study translates the advanced deep learning technique to clinical practice where AMT-PET is currently unavailable. It was found that increased AMT-learned MRI tumor volume can be prognostic for overall survival, independent of specific multi-modal MR protocols (hardware and sequence parameters etc.). This finding provides preliminary evidence to support that an end-to-end deep learning of multi-modal MRI data can be effective and feasible for survival prediction of glioblastoma patient without the need of any additional measurement or procedure such as AMT-PET.Conclusion
Systematic investigation of the proposed U-net approach may improve presurgical evaluation in glioblastoma by supplementing conventional multi-modal MRI to approximate glioblastoma volume with high amino acid uptake.Acknowledgements
This work was supported by a grant from the National Institute of Health, R01 CA123451 (C.J and S.M).References
1. Juhász C, Muzik O, Chugani DC, Chugani HT, Sood S, Chakraborty PK, Barger GR, Mittal S. Differential kinetics of α-[¹¹C]methyl-L-tryptophan on PET in low-grade brain tumors. J Neurooncol. 2011;102(3):409-15. 2. Juhász C, Chugani DC, Barger GR, Kupsky WJ, Chakraborty PK, Muzik O, Mittal S. Quantitative PET imaging of tryptophan accumulation in gliomas and remote cortex: correlation with tumor proliferative activity. Clin Nucl Med. 2012;37(9):838-42. 3. Kamson DO, Mittal S, Robinette NL, Muzik O, Kupsky WJ, Barger GR, Juhász C. Increased tryptophan uptake on PET has strong independent prognostic value in patients with a previously treated high-grade glioma. Neuro Oncol. 2014;16(10):1373-83. 4. Bosnyák E, Michelhaugh SK, Klinger NV, Kamson DO, Barger GR, Mittal S, Juhász C. Prognostic Molecular and Imaging Biomarkers in Primary Glioblastoma. Clin Nucl Med. 2017;42(5):341-347. 5. Blanc-Durand P, Van Der Gucht A, Schaefer N, Itti E, Prior JO. Automatic lesion detection and segmentation of 18F-FET PET in gliomas: A full 3D U-Net convolutional neural network study. PLoS One. 2018;13(4):e0195798. doi: 10.1371/journal.pone.0195798. 6. Ronneberger O., Fischer P., Brox T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In: Navab N., Hornegger J., Wells W., Frangi A. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. MICCAI 2015. Lecture Notes in Computer Science, 9351:234-41. Springer, Cham.Figures
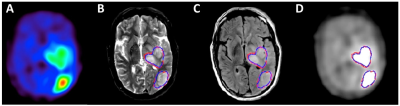
Figure 1. (A)
AMT-PET of a left hemispheric glioblastoma with two lesions showing high
uptake. Tumor volumes identified by U-Net fully learned two multi-modal MRI
features, T2 (B) and FLAIR (C). Blue and red contours represent
U-Net-determined glioma vs. ground truth (i.e., increased uptake above 1.65 ratio
of tumor and normal cortex as defined on AMT-PET [D]).
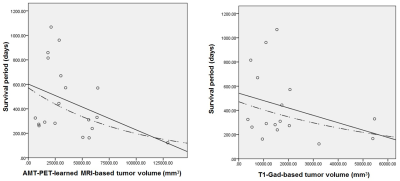
Figure 2. Correlation between pre-treatment
tumor volume and overall survival in 19 patients. The PET-learned MRI volumes
showed a significant negative (exponential) correlation (R2=0.24,
p=0.03), while the correlation with the T1-Gad contrast-enhancing tumor volume
was not significant (p>0.1 in both linear and exponential correlations).
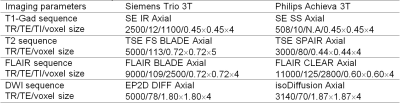
Table 1. Multi-modal MRI protocols.
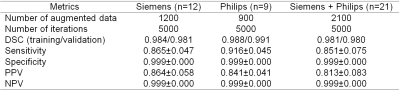
Table 2. Overall performance of individual
U-Net system to detect AMT-PET tumor volume from multi-modal MRI data.