2345
Genomically and Radiographically Adjusted Dose (GRAD) Framework for Biologically Adaptive MR-guided Radiotherapy1Biophysics, Medical College of Wisconsin, Milwaukee, WI, United States, 2Radiation Oncology, Medical College of Wisconsin, Milwaukee, WI, United States, 3Radiology, Medical College of Wisconsin, Milwaukee, WI, United States
Synopsis
Tumor burden, tumor proliferation, and tumor hypoxia, all of which vary in space and time, are evidence-based contributors of radiotherapy failure. In addition, it has been demonstrated that gene expression can influence radiosensitivity. We demonstrate here the initial feasibility of a framework to incorporate genomic and radiographic information to derive patient-specific, voxelwise radiation dose prescription maps for use in a biologically adaptive MR-guided radiotherapy (BAMRgRT) strategy.
Introduction
The objective of conventional radiotherapy is to deliver uniform doses to tumor targets while minimizing dose to surrounding organs at risk. However, tumor burden, tumor proliferation, and tumor hypoxia, evidence-based contributors of radiotherapy failure, can vary over space and time. Furthermore, it has recently been demonstrated that gene expression can influence radiosensitivity1. We demonstrate here a framework to incorporate genomic and radiographic information to derive patient-specific, voxelwise radiation dose prescription maps for use in a biologically adaptive MR-guided radiotherapy (BAMRgRT) strategy.Methods
The feasibility of genomically and radiographically adjusted dose (GRAD) for BAMRgRT was tested in two brain tumor patients. A quantitative MRI (qMRI) panel consisting of i) multiple flip angle T1 mapping (3, 6, 10, 20, 30 degrees), ii) accelerated CPMG T2 mapping (11-200 msec), iii) IVIM DWI (10 b-values), and iv) pharmacokinetic DCE-MRI was acquired in each patient. To assess the absolute bias and short-term repeatability of qMRI parameters, calibration experiments were performed using the Eurospin TO5 and QIBA diffusion phantoms on an Elekta 1.5T MR-Linac and Siemens 1.5T and 3.0T MR simulators. A reference genomic-adjusted radiation dose (GARD) score was calculated for a conventional 2 Gy per fraction treatment using a population radiosensitivity index (RSI) of 0.2 and beta of 0.0671. The GARD score was then used to determine the required dose for a patient-specific RSI of 0.25. For each patient, gross tumor and high-risk tumor volumes were manually segmented on the qMRI parameter maps. A voxelwise 3D dose prescription map was then calculated in MIM Maestro utilizing the linear-quadratic model extended to incorporate GARD. An inverted copy of the calculated 3D dose prescription map was also constructed for use in treatment planning. Synthetic CT images, generated from a high resolution 3D T1 FLASH image of each patient using a conditional GAN2, and inverted 3D dose prescription maps were loaded into the Elekta Monaco radiation treatment planning system (RTPS). Volumetric modulated arc therapy (VMAT) plans (3mm dose grid, 1% statistical uncertainty per calculation) were calculated with dose painting by numbers (DPBN)3, facilitated by using the inverted dose prescription map as a bias dose during plan optimization. Finally, sample VMAT plans were calculated in phantom to determine the modulation transfer function (MTF) of Monaco, in order to investigate the degree to which the RTPS is able to modulate dose of proximal clusters of high-risk tumor cells.Results
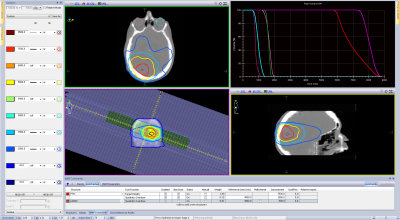
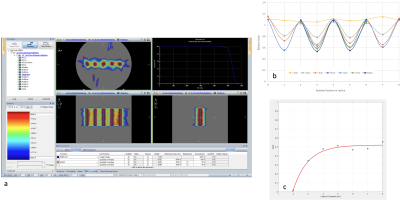
Figure 1 displays a qMRI panel for one representative patient. Figure 2 displays results of calibration experiments for T1, T2, and apparent diffusion coefficient parameter maps. The accuracy and short-term repeatability of the MR-Linac was comparable to the MR simulators studied. Figure 3 displays sample forward and inverted 3D dose prescription maps overlaid onto a synthetic CT calculated for one of the patients studied. The tumor regions were manually segmented on the qMR images. The two dose levels of 58.6 Gy and 75.3 Gy were obtained using the linear-quadratic model extended to incorporate GARD. Figure 4 displays a DPBN radiation treatment plan calculated in Monaco using the inverted dose prescription map as a bias dose. Both PTVs were able to achieve over 95% coverage and the global max dose of the plan was approximately 16%. Figure 5 displays the MTF results of the Monaco RTPS. For 1 cm diameter targets, about a 40% modulation of dose was achievable with a 3.5 cm lattice spacing. The dose modulation reached an asymptote of about 50% beyond a lattice spacing of about 5 cm.Discussion
These initial results demonstrate the feasibility of a framework to genomically and radiographically adjust radiotherapy doses for BAMRgRT. The inverted dose prescription maps facilitate DPBN on commercial RTPS, which normally do not support voxelwise objective functions. Once a treatment plan is obtained, the forward dose prescription map can be employed to evaluate the calculated plan quality or expected tumor control probability. As shown by the MTF study, the Monaco RTPS has limitations in the extent to which dose can be modulated. This may have implications on DPBN of small, high-risk tumor cell clusters demanding rapid variations in dose. Future work will include development of machine learning methods to automatically segment gross and high-risk tumor regions from qMRI parameter maps.Conclusion
A feasible framework has been created to incorporate genomic and radiographic information to derive patient-specific radiation dose prescription maps for use in a biologically adaptive MR-guided radiotherapy (BAMRgRT) strategy.Acknowledgements
Advancing a Healthier Wisconsin (5520223/5520231)References
1. J. G. Scott et al., “A genome-based model for adjusting radiotherapy dose (GARD): a retrospective, cohort-based study,” Lancet Oncol., vol. 18, no. 2, pp. 202–211, 2017.
2. P. Isola, J. Y. Zhu, T. Zhou, and A. A. Efros, “Image-to-image translation with conditional adversarial networks,” Proc. - 30th IEEE Conf. Comput. Vis. Pattern Recognition, CVPR 2017, vol. 2017–Janua, pp. 5967–5976, 2017.
3. S. M. Bentzen, “Theragnostic imaging for radiation oncology: Dose-painting by numbers,” Lancet Oncol., vol. 6, no. 2, pp. 112–117, 2005.
Figures
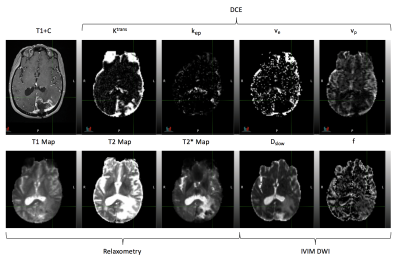
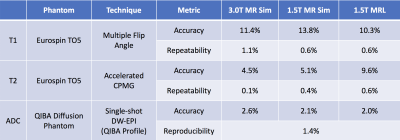
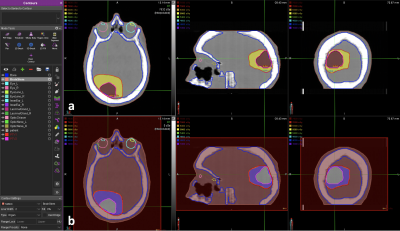
Figure 2: Forward (a) and inverted (b) patient-specific GRAD dose prescription maps overlaid on synthetic CT images. Gross and high-risk tumor regions were manually segmented using qMRI parameter maps. Dose levels were calculated using the linear-quadratic model extended to incorporate GARD.