0475
Reconstructing Lesions not seen during training using a Recurrent Inference MachineBob van Hoek1, Kai Lønning2,3, Hanneke Hulst4, Frans Vos1,5, and Matthan Caan6
1Delft University of Technology, Delft, Netherlands, 2Radiotherapy, Netherlands Cancer Institute, Amsterdam, Netherlands, 3Spinoza Centre for Neuroimaging, Amsterdam, Netherlands, 4Anatomy and Neurosciences, VU University Medical Center, Amsterdam, Netherlands, 5Radiology, Academic Medical Center, Amsterdam, Netherlands, 6Biomedical Engineering and Physics, Academic Medical Center, Amsterdam, Netherlands
Synopsis
Deep learning can accelerate MRI beyond what is currently possible. Broad clinical application requires generalizability to multiple contrasts, acceleration levels and pathologies. Here we explore how a Recurrent Inference Machine trained on healthy volunteer T1-weighted brain images performs in such a situation, by reconstructing FLAIR images with white matter lesions, in simulation and prospectively undersampled patient data. Lesion contrast is maintained up to 6x acceleration and higher than in compressed sensing (CS) reconstruction, and all lesions are retained compared to CS.
Introduction
Deep learning based MRI-reconstruction is showing a great promise in speeding up both acquisition time by sparse sampling as well as reconstruction time thanks to fast inference times using pre-trained networks on fast GPU-cards. In clinical practice, a wide variety of protocols is used, with a range of sequence parameters and designs that will be challenging to include in a training set. Most work to date is experimenting within the close measurement conditions of the training data, including acceleration level and MR-contrast. Here, promising results are reported, e.g. in detecting subtle pathologies in the knee joint$$$^1$$$. Our aim is to explore the unseen landscape of sequence parameters and pathologies, in a case study on reconstructing brain lesions in FLAIR data novel to a trained network.Methods
The experimental overview and network architecture are depicted in Figs.1&2. Instead of learning a direct mapping from the sparsely sampled image to the full signal, Recurrent Inference Machines$$$^2$$$ (RIM) learn to reconstruct the signal in an iterative procedure similar to compressed sensing, where the goal is to optimize the maximum a posteriori estimate. To this aim it employs a recurrent architecture, where each step improves on the previous step’s reconstruction estimate by using as input the gradient of the log-likelihood distribution, as given under the assumption of normally distributed noise, along with the model’s current reconstruction state, which should enable it to estimate the gradient of the log-prior distribution. Due to their recurrent architecture, RIMs swap model parameters for internal states, moving dependency away from the training data and onto the input data instead. This prevents overfitting and enables the reconstruction of features that are not necessarily present in the training set, i.e. lesions after only being exposed to healthy volunteers.We trained a model of approximately 100k parameters, using the mean square error averaged over the model’s 8 reconstruction estimates as a loss function. Raw data of 12 participants was acquired on a Philips 3T scanner (Philips, The Netherlands) with a 32-channel head coil. For training, fully sampled 3D T1-weighted MPRAGE data at 1mm was acquired in 10 healthy volunteers (scanning time 10 minutes). For testing purposes, an accelerated 3D FLAIR scan of a single healthy volunteer was obtained at 1.1mm, with a sparse sampling pattern 6x as implemented by the vendor (scanning time 5 minutes). For validation, an identical 3D FLAIR was used to scan one patient with a mild white matter lesion load. Coil sensitivity maps were computed from the fully sampled center in all scans with calibration size 15 with ESPIRiT auto calibration$$$^3$$$.Two lesions were simulated in one slice of the healthy volunteer 3D-FLAIR testing dataset, initially reconstructed using compressed sensing (CS) for this purpose. Using manual annotation, the signal magnitude was augmented to create local hyperintensities (Fig. 1).The signal phase was not adjusted, assuming negligible phase shift. The resulting image was multiplied by coil sensitivity maps, Fourier transformed and subsampled with factors 4-10x. CS reconstruction was performed using the Berkeley Advanced Reconstruction toolbox, with an l1-norm sparsity term using the wavelet transform, and optimized regularization (r=0.005). RIM-reconstructions were performed using the network trained on T1-weighted data. In the reconstructed images, lesion contrast was computed as $$$(I_{peak}-I_{ref})/I_{ref}$$$, with $$$I_{peak}$$$ the peak intensity within the lesion and $$$I_{ref}$$$ the mean intensity in an adjacent reference intensity region.Results
The RIM was trained until convergence in 1 day on a single GPU-card. T1-weighted images could be reconstructed at high acceleration levels (Fig.1). Inference times for single slices were approximately 0.1s. Figure 3 shows that at 30% intensity, the lesion is slightly more apparent in the RIM-reconstruction than in the CS-reconstruction. Figure 4 quantifies lesion contrast, showing that at 4x- and 6x-acceleration the RIM maintains contrast while at 8x- and 10x-acceleration more variable contrast for the two lesions is seen. CS underperforms at all acceleration levels. Figure 5 illustrates that in prospectively undersampled patient data, the T1w-trained RIM does not underperform on reconstructing WM lesions compared to CS.Discussion and Conclusion
The RIM can reconstruct FLAIR images with small lesions that were not presented during training. Our lesion model of multiplying by small Gaussians corresponds to convolutions with large Gaussians in Fourier domain. Thus, a large extent of k-space, far beyond the sub-sampling range, contributes to the lesion signal. Future work is needed to assess how other deep learning methods that operate solely in image domain perform compared to the RIM.Acknowledgements
No acknowledgement found.References
- Hammernik K, Klatzer T, Kobler E, Recht MP, Sodickson DK, Pock T, Knoll F (2018): Learning a variational network for reconstruction of accelerated MRI data. Magn Reson Med 79:3055–3071.
- Putzky P, Welling M. Recurrent Inference Machines for Solving Inverse Problems. arXiv:1706.04008, 2017.
- Uecker M, Lai P, Murphy MJ, Virtue P, Elad M, Pauly JM, Vasanawala SS, Lustig M. ESPIRiT — An Eigenvalue Approach to Autocalibrating Parallel MRI : Where SENSE Meets GRAPPA. Magn. Reson. Med. 2014;1001:990–1001.
Figures
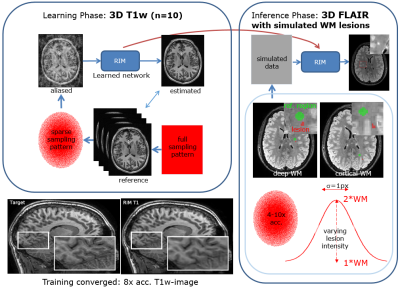
Figure 1. A Recurrent Inference Machine (RIM) was trained on 10 non-accelerated T1-weighted scans. Training successfully converged, illustrated by a reconstructed T1w-scan (lower-left). After that, FLAIR data with simulated lesions were reconstructed using the trained RIM. Lesions had varying intensity achieved by multiplying by values (1+a) with a=[0…100%], and are bandwidth-limited by convolving with a Gaussian kernel with $$$\sigma$$$=1px. Data were accelerated over a range of 4-10x.
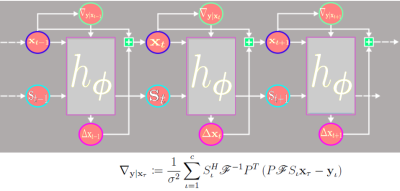
Figure 2. The Recurrent Inference Machine (RIM) unrolled in time. The network parameters are shared over time steps. Based on the current estimate of the image $$$x_t$$$, the log-likelihood gradient $$$\nabla_{y|x_t}$$$ is computed based on the forward model with $$$c$$$ coil sensitivities $$$S$$$ and undersampling pattern $$$P$$$. The network $$$h_\phi$$$ contains two GRU’s embedded in three CNNs and two hidden states $$$s$$$. The image update $$$\Delta x_t$$$ is add to obtain the new estimated image.
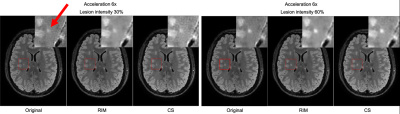
Figure 3. Illustration of a simulated lesion at intensities of 30% and 60% in the original FLAIR slice, with reconstructions using the Recurrent Inference Machine (RIM) and Compressed Sensing (CS). The lesion appears as a small punctate object, adjacent to cortical GM.
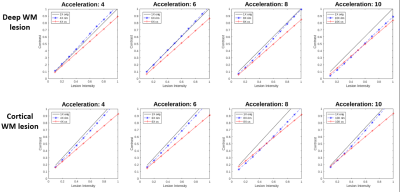
Figure 4. Lesion contrast as a function of simulated lesion intensity for the original scan (1x orig), the Recurrent Inference Machine (RIM) and Compressed Sensing (CS) for two modelled lesions in deep and cortical WM.
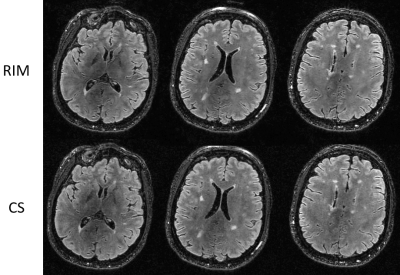
Figure 5. Reconstructed images of 6x prospectively undersampled data of a patient with a mild white matter lesion load. Note that all lesions remain visible in the RIM-reconstruction compared to the CS-reconstruction.