5457
Novel multi-slab GRASE sequences for fMRI. Comparison with EPI.1Radiology, Center for Magnetic Resonance Research, University of Minnesota, Minneapolis, MN, United States
Synopsis
T2 weighted fMRI has been shown to provide higher spatial specificity than commonly used T2* contrast. However, 2D-spin-echo-EPI method, which is the most commonly employed approach, suffers from a low functional mapping contrast. Versions of single-shot 3D-GRASE sequence provide improvements in fMRI contrast, but suffer limitations particularly at ultrahigh magnetic fields due to shorter T2. To solve this problem, a new multi-slab version of the 3D-GRASE sequence with and without the capability of inner-volume selection, abbreviated as ivmsGRASE and msGRASE, respectively, are proposed. The theoretical and practical considerations of these methods in comparison with spin-echo-EPI for fMRI application are presented.
Introduction
Comparatively, T2 weighted spin-echo contrast fMRI has been proved to yield higher spatial specificity than T2* contrast, due to its selectivity for smaller blood vessels involved in the neuro-vascular coupling, which provides the functional mapping signals in fMRI1. However, conventional 2D-spin-echo-EPI (seEPI) suffers from a low BOLD functional contrast-to-noise ratio (fCNR)2. To overcome this obstacle, single-shot 3D-GRASE combining fast spin echo acquisition scheme with EPI readout has been implemented3. The multiple refocusing pulses employed after a single excitation increase signal-to-noise ratio (SNR) and fCNR due to the Fourier averaging. By using slab-selective excitation and refocusing pulses in orthogonal axes an inner-volume can be imaged4. The inner-volume selection allows shorter echo train, which alleviates blurring effect on the phase-encoding direction. This blurring remains a major limitation, especially at ultrahigh magnetic field due to the significantly shorter T2. To solve this problem, we propose a single-shot multi-slab 3D-GRASE sequences with and without the capability of inner-volume selection, respectively abbreviated as ivmsGRASE and msGRASE. Below we present the theoretical and practical considerations of these methods in comparison with seEPI for fMRI application.Methods
Fig.1 schematically presents the discussed pulse sequences. All three sequences use the similar seEPI readout scheme. The difference lies only in the way the third dimension is encoded. EPI uses stacked slices while GRASE uses slabs, consisting of spin echoes (encoding in plane defined by the readout and phase encoding) with a second phase encodings in the third dimension. These three sequences were compared under condition of the same spatial resolution, matrix size (unless specified for the inner-volume case), and repetition time between single-shot fMRI image collections. The noise was assumed to be the same for all sequences and was not considered. Similarity of these sequences allows simplified SNR comparison based on differences of the three components, such as: SNR~“Excitation_efficiency”x“T2_relaxation”x“Acquisition_efficiency”. These components are presented in detail in Table1. For discussion below we use a normalized SNRNmsGRASE and SNRNivmsGRASE calculated as ratio of SNR of GRASE sequences to SNR of seEPI. Experiment: The single-shot msGRASE sequences where implemented and tested on an in vivo rat brain on a Varian 16.4T animal scanner with single loop 2.5cm surface coil. Resting-state fMRI was further acquired with conventional seEPI and proposed GRASE sequences.Results
Higher values of npe2 benefits SNR, however to avoid blurring we must keep npe2<πT2/2τ and reach a target resolution by increasing the number of slabs. The inner-volume case uses smaller npe and allows increased npe2. The “Excitation_efficiency” depends on effective TR and is lowest for ivmsGRASE. Accelerating recovery of longitudinal magnetization due to blood in-flow effect as well as longer effective in Carr-Purcell echo train with decreased echo-time in brain tissue5,6 highly improve SNR for ivmsGRASE. The results of theoretical predictions for the normalized SNRs are presented in Table2. The experimental data presented at Fig.2 in general agreed with theoretical predictions. We found highly shortened magnetization recovery time in the brain tissue (~0.5s), which is about 4 times lower than T1 of rat brain tissue at 16.4T (~2.3s)7. This expected in-flow (perfusion) effect arising from the use of a small surface coil allowed us to increase flip angles considerably, which, as a result, improved SNR, especially for ivmsGRASE images.Discussion
The theoretical and practical considerations based on fMRI experiments show that single-shot multi-slab 3D-GRASE is a good alternative to seEPI for functional studies. The multi-slab inner-volume capability is even more practical, especially at ultrahigh field with shortened T2 relaxation. Imaging with surface coils is very beneficial in this case due to blood in-flow (perfusion) signal enhancement. However, influence of this effect on BOLD contrast is yet to be determined. One considerable complication in multi-slab acquisition is the striped artifacts along the slab selection due to non-ideal excitation profile. However it was shown that uneven profiles could be compensated almost lossless during image post-processing8. Even without compensation, these artifacts have low impact on fMRI results. Additionally, the performance of these sequences could be enhanced by utilizing partial Fourier9 and multi-band accelerations10.Conclusion
A novel multi-slab single-shot 3D-GRASE sequences have been presented and compared with conventional spin-echo EPI at high field for high resolution fMRI study. In in-vivo application of the presented technique shows high flexibility and considerable better SNR and T2 contrast compared to conventional spin-echo EPI.Acknowledgements
This study was supported by NIH grants R01 MH111447 and MH111413; P41 EB015894; P30 NS07640.References
1. Yacoub E, Shmuel A, Logothetis N, Ugurbil K. Robust detection of ocular dominance columns in humans using Hahn Spin Echo BOLD functional MRI at 7 Tesla. NeuroImage 2007;37(4):1161-77. 2. Ugurbil K. What is feasible with imaging human brain function and connectivity using functional magnetic resonance imaging. Philosophical Transactions of the Royal Society B: Biological Sciences 2016;371(1705):20150361. 3. Oshio K, Feinberg DA. Single-Shot GRASE Imaging without Fast Gradients. Magn Reson Med 1992;26(2):355-360. 4. Feinberg DA, Harel N, Ramanna S, Ugurbil K, Yacoub E. Sub-millimeter single-shot 3D GRASE with inner volume selection for T2 weighted fMRI applications at 7 Tesla. In: 16th Annual Meeting International Society for Magnetic Resonance in Medicine. Toronto, ON; 2008. p. 2373. 5. Jensen JH, Chandra R, Yu H. Quantitative model for the interecho time dependence of the CPMG relaxation rate in iron-rich gray matter. Magn Reson Med 2001;46(1):159-65. 6. Michaeli S, Garwood M, Zhu X-H, DelaBarre L, Andersen P, et al. Proton T2 relaxation study of water, N-acetylaspartate, and creatine in human brain using Hahn and Carr-Purcell spin echoes at 4T and 7T. Magn Reson Med 2002;47(4):629-633. 7. Pohmann R, Shajan G, Balla DZ. Contrast at high field: Relaxation times, magnetization transfer and phase in the rat brain at 16.4 T. Magn Reson Med 2011;66(6):1572-1581. 8. Wu W, Koopmans PJ, Frost R, Miller KL. Reducing slab boundary artifacts in three-dimensional multislab diffusion MRI using nonlinear inversion for slab profile encoding (NPEN). Magn Reson Med 2016;76(4):1183-1195. 9. Feinberg DA, Hale JD, Watts JC, Kaufman L, Mark A. Halving MR imaging time by conjugation: demonstration at 3.5 kG. Radiology 1986;161(2):527-531. 10. Feinberg DA, Setsompop K. Ultra-Fast MRI of the Human Brain with Simultaneous Multi-Slice Imaging. Journal of magnetic resonance (San Diego, Calif : 1997) 2013;229:90-100. 11. Ernst RR, Anderson WA. Application of Fourier transform spectroscopy to magnetic resonance. Rev Sci Instrum 1966;37:93-102. 12. Ernst RR, Bodenhausen G, Wokaun A. Principles of Nuclear Magnetic Resonance in One and Two Dimensions. 5-th ed. Oxford: Clarendon Press; 1997. 518 p.Figures
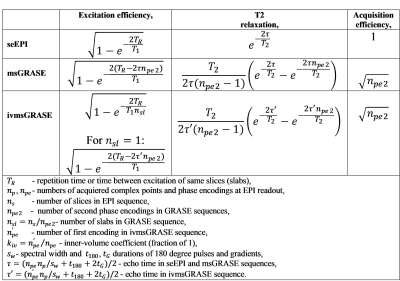
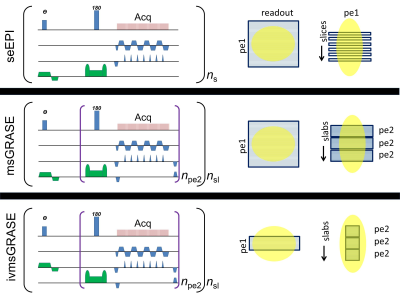

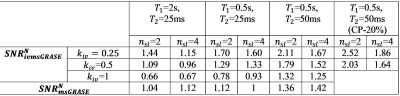
Fig.2 (A) Anatomical images using multi-slice fast spin echo (FSE);
(B) Conventional SE-EPI images with resolution of 0.375x0.375x0.5 mm3 ;
(C) ivmsGRASE images with the same resolution as seEPI images. Based on the ICA analysis, a resting state network on somatosensory area is clearly shown.