3976
Optimal phased-array signal combination from separate coil elements for GABA quantification at 7T1Erwin L. Hahn Institute for MRI, University of Duisburg-Essen, Essen, Germany, 2Donders Institute for Brain, Cognition and Behavior, Radboud University, Nijmegen, Netherlands
Synopsis
Hitherto, signal combination strategies from separate coil elements have been evaluated on the basis of spectral SNR improvement. This study compared various combination methods for two representative GABA measurement techniques: GABA editing and short echo time acquisitions, and investigated which signal combination method is optimal in terms of GABA quantification using LCModel.
Introduction
To take maximum advantage of phased array coil acquisition, optimal combination strategies are required. The widely used signal weighted combination1 is necessarily not optimal for low SNR metabolites, such as GABA, which is present in circa one millimolar concentration. Previously, signal combination strategies from separate coil elements have been evaluated on the basis of spectral SNR improvement. In this study, we compared various combination methods for two representative GABA measurement techniques: GABA editing and short echo time (TE) acquisitions, and investigated which signal combination method is optimal in terms of GABA quantification using LCModel2, which is one of the most popular spectral quantification tools.Methods
MEGA-sLASER3 and short TE sLASER4 sequences acquired single voxel spectroscopy data from occipital and parietal cortex of 15 healthy volunteers (4M / 11F; 29.6 ± 4.69YO) with following parameters: 7T system (Siemens, Erlangen), 32ch head coil (Nova medical, Wilmington, MA), 8cm3 isotropic voxel, MEGA-sLASER: TE/TR/NEX = 80ms/4500ms/64, and sLASER: TE/TR/NEX = 38ms/4500ms/64. A 3D MPRAGE was acquired as an anatomical reference. B0 shimming was performed by FAST MAP5.
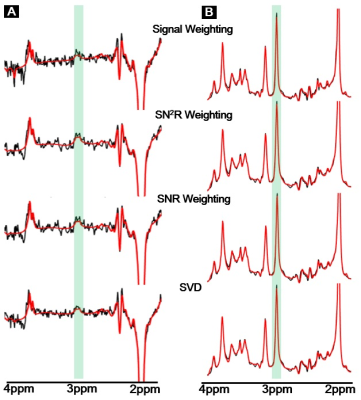
First, the spectra from each coil element was realigned by 0thorder phase-correction in the time domain, and combined by four methods of signal combination: 1) a signal weighting method performed by multiplying coil sensitivity by each raw signal, here the coil weighting was selected as the highest amplitude of the time domain signal before FT: this signal weighting approach is used by the manufacturer; 2) SNR weighting, where the S/N of the target peak in the frequency domain after FT provides the weighting coefficient6; 3) SN2R weighting, the S/N2 of the target in the frequency domain is used as the weighting coefficient7, and 4) Singular value decomposition (SVD), the principal signal component was extracted from noise by the computed noise covariance matrix, which follows a multivariate normal distribution8. This matrix was calculated by white noise area of the time domain signal. For frequency domain weightings, 3ppm peak was used as the target peak.
All signal combination steps were processed using custom written script in Matlab (ver. 2016b, Mathwork, Natick, MA). SNR of individual spectra from different combination methods was estimated by the 3ppm peak: edited GABA peak for MEGA-sLASER and creatine peak for sLASER. LCModel (ver. 6.3, Provencher, Ontario) quantified 19 Metabolites’ (listed in Figure2) concentrations with Cramer Rao Lower Bound (CRLB), which is an estimated fitting error.
Results
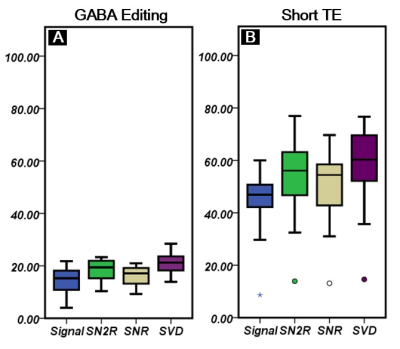
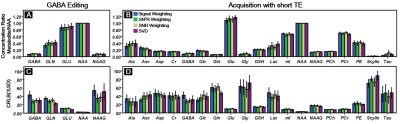
Figure1A (GABA editing) and 1B (Short TE) show combined signals by four different combination methods with a corresponding fitting line by LCModel. Figure2 shows SNR comparison of the 3ppm peak in GABA editing(A) and short TE(B)approaches. In both cases, we could find a relatively high SNR at SVD, and a relatively low SNR at signal weighting as compared with other two SNRs by SNR and SN2R weightings were similar. Figure3 compares metabolic concentration ratios (A and B), which were normalized by NAA, and CRLB (C and D) between weighting methods of both acquisition approaches. For GABA editing, we could not find a notable concentration difference between weightings. However, SN2R method increased the fitting reliability in comparison with other methods. For the short TE acquisition, there was no apparent difference in metabolic concentration and CRLB between weighting methods for high concentrated metabolites such as NAA, Cr and PCh. Whereas low concentrated metabolites such as Tau, Lac, and Ala showed wider variations in concentration ratios and CRLB than those of high concentrated metabolites. SN2R weighting also showed a relatively low CRLB value than those of other schemes at GABA in the short TE acquisition.Discussion
This study has compared the spectral quality and the spectral fitting quality between different coil signal combination strategies. The SVD method accomplished a significant SNR improvement by eliminating noise effectively. Nevertheless, the SN2R weighting provided reliable GABA quantification by decreasing spectral fitting error. As the 3ppm peak was used for the weighting coefficient in the frequency domain weighting methods, optimized 3ppm peak enabled accurate GABA quantification by spectral fitting. Judging from the spectral fitting quality, SN2R weighting is an optimal coil combination method for GABA quantification. In addition, optimized SN2R weighted combination does not require a dominant signal in the time domain, therefore, we can use strong water suppression, which can eliminate baseline distortion and spurious signals, and lead to the reliable and consistent detection of low concentration metabolites.Acknowledgements
This work was funded by the Helmholtz Alliance ICEMED – Imaging and Curing Environmental Metabolic Diseases, through the Initiative and Networking Fund of the Helmholtz Association.References
1. Roemer, Peter B., et al. "The NMR phased array." Magnetic resonance in medicine 16.2 (1990): 192-225.
2. Provencher, Stephen W. "Estimation of metabolite concentrations from localized in vivo proton NMR spectra." Magnetic resonance in medicine 30.6 (1993): 672-679.
3. Scheenen, Tom WJ, et al. "Short echo time 1H‐MRSI of the human brain at 3T with minimal chemical shift displacement errors using adiabatic refocusing pulses." Magnetic resonance in medicine 59.1 (2008): 1-6.
4. Andreychenko, Anna, et al. "Efficient spectral editing at 7 T: GABA detection with MEGA‐sLASER." Magnetic resonance in medicine 68.4 (2012): 1018-1025.
5. Gruetter, Rolf, and Ivan Tkáč. "Field mapping without reference scan using asymmetric echo‐planar techniques." Magnetic resonance in medicine 43.2 (2000): 319-323.
6. Wald, Lawrence L., et al. "Proton spectroscopic imaging of the human brain using phased array detectors." Magnetic resonance in medicine 34.3 (1995): 440-445.
7. Hall, Emma L., et al. "Methodology for improved detection of low concentration metabolites in MRS: optimised combination of signals from multi-element coil arrays." Neuroimage 86 (2014): 35-42.
8. Erdogmus, Deniz, et al. "Image construction methods for phased array magnetic resonance imaging." Journal of Magnetic Resonance Imaging 20.2 (2004): 306-314.
Figures