3529
Joint Reconstruction of Images with Different Temporal Basis in Carotid Vessel Wall Imaging1Biomedical Imaging Research Institute, Cedars-Sinai Medical Center, Los Angeles, CA, United States, 2Department of Bioengineering, University of California, Los Angeles, Los Angeles, CA, United States, 3Department of Medicine, University of California, Los Angeles, Los Angeles, CA, United States
Synopsis
We recently proposed two techniques, qMATCH and DCE, based on Low Rank Tensor (LRT) framework. qMATCH is a single 8-min scan for carotid T1 T2 mapping and LRT DCE is a 10-min scan evaluating inflammatory status of carotid atherosclerosis. The LRT DCE has many advantages over conventional DCE protocols, but has a scan time longer than typical 5-6 minutes. In this work, we proposed a new protocol combining qMATCH and 5-min DCE. In vivo studies have demonstrated the feasibility of the joint reconstruction. Results of joint reconstruction showed improved image quality with shortened scan time.
Introduction
Imaging methods to evaluate vulnerable plaques for carotid atherosclerosis are of substantial interest1-3. Recently, we proposed two novel protocols, qMATCH and 3D DCE for quantitatively evaluation of carotid plaque using low-rank tensor (LRT) imaging4-6. qMATCH involves a single 8-min scan performing 3D T1-T2 mapping, allowing detection of plaque composition. 3D LRT DCE involves 10-min scan with 0.7mm isotropic spatial resolution, 0.59s temporal resolution and fast T1 mapping for assessment of inflammatory status. 3D DCE has many advantages over previous DCE protocols, but has a scan time longer than the typical 5-6 minutes. Because qMATCH and 3D LRT DCE images of the same anatomy are correlated, joint reconstruction of both sets of images has the potential to increase the image quality for both with shorter total scan time. In this work, we proposed a new protocol combining qMATCH and 5-min DCE, investigating the effect on the image quality of DCE.Methods
Sequence design and sampling pattern: As described previously5,6, qMATCH part employed 3D flow-compensated spoiled gradient echo readout (FLASH) with variable-duration T2-IR preparations to generate T1 and T2 contrast. The DCE sequence implemented the same FLASH readout following SR preparations. Cartesian acquisition with randomized reordering in ky and kz directions was implemented according to a variable-density Gaussian distribution. The center k-space line was collected every eighth readout to serve as LRT training data4.
Joint reconstruction: qMATCH images form a 3-way tensor $$$\mathscr{A}_1$$$ with spatial dimension $$$\mathbf{r}=(x,y,z)$$$, an IR dimension $$$T_I$$$, and a T2prep-duration dimension $$$T_E$$$. DCE images form a 3-way tensor $$$\mathscr{A}_2$$$ with a spatial dimension $$$\mathbf{r}$$$, an SR dimension $$$\tau$$$, and a DCE dimension t. Due to the strong correlation between images with different contrasts, each tensor is low-rank with a shared spatial subspace and can be factorized as $$$\mathbf{A}_{1,(1)}=\mathbf{U\Phi}_1$$$ and $$$\mathbf{A}_{2,(1)}=\mathbf{U\Phi}_2$$$, where the rows of $$$\mathbf{\Phi}_1$$$ and $$$\mathbf{\Phi}_2$$$ span temporal tensor subspaces for qMATCH and DCE, respectively. In the joint recon, $$$\mathbf{\Phi}=[\mathbf{\Phi}_1,\mathbf{\Phi}_2]$$$ are determined together from subspace training data $$$\mathbf{D}_{\rm{tr}}=[\mathbf{D}_{\rm{tr,1}},\mathbf{D}_{\rm{tr,2}}]$$$. $$$\mathbf{U}$$$, the spatial coefficients for both qMATCH and DCE, is then recovered by fitting $$$\mathbf{\Phi}$$$ to the remainder of the sparsely sampled data.
Motion removal: Abrupt patient motion is a major challenge for vessel wall scan due to the small anatomical structure. In the LRT framework, abrupt-motion frames are less correlation to the remainder of the images, corresponding to a higher residual after LRT estimation. The residual $$$\boldsymbol{r}_{res}$$$ is calculated by: $$\boldsymbol{r}_{res}=\sum\nolimits_{i}|(\mathbf{D}_{\rm{tr}}-\mathbf{D}_{\rm{tr}}\mathbf{\Phi}^\dagger\mathbf{\Phi})_{ij}|^2,$$Where $$$\mathbf{D}_{\rm{tr}}\in\mathbb{C}^{N_r\times N_t}$$$ is the matrix form of subspace training data. $$$N_r$$$ and $$$N_t$$$ represent the total number of voxels and time points, respectively. High-residual time points were removed and recovered by LRT framework during the recovery of subspace training data inherently4.
Imaging protocol: All data were acquired on a 3T commercial MR scanner (MAGNETOM Verio, Siemens Healthineers, Erlangen, Germany). Healthy subjects (n=2) were scanned with the following parameters: coronal orientation, spatial resolution=0.7mm isotropic, FOV=150x150x26mm3, $$$\alpha =8^\circ$$$, echo spacing=11ms. For qMATCH part, TR=2360ms, T2-prep duration=20/30/40/50/60/70ms, scan duration=8min. For DCE part, TR/temporal resolution=590ms, scan duration = 5min.
Data analysis: We reconstructed the acquired data in two ways. One is the described joint reconstruction, the other is solo reconstruction of the 5-min DCE data only6. In the analysis, we compared the SNR and sharpness of DCE images from these two reconstruction methods.
Results
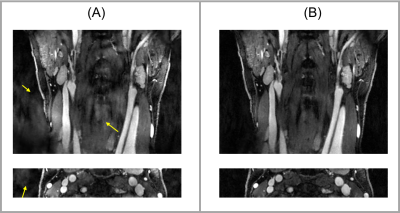
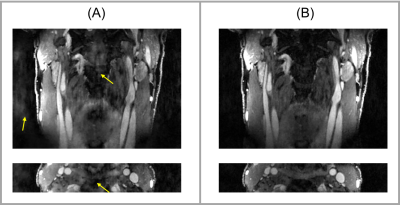
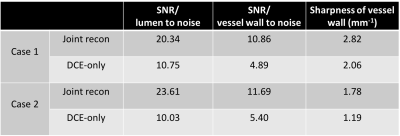
Figures 1 and 2 displays example images for joint reconstruction and DCE-only reconstruction at peak enhancement. With joint reconstruction, the artifacts were alleviated and the delineation was sharper compared to DCE-only recon. Table 1 lists the SNR of lumen and wall area and sharpness of the vessel wall for both the joint reconstruction and DCE-only reconstruction.Discussion
This study showed the feasibility of joint reconstruction of qMATCH and 5-min DCE images. The results illustrated that the combination of qMATCH and DCE increases the image quality of DCE portion. We will further investigate the effect of the joint recon on the qMATCH portion.Conclusion
In this work, we demonstrated the feasibility of joint reconstruction of qMATCH and DCE in carotid vessel wall imaging. Results showed that the combination of qMATCH and DCE can substantially increase the image quality of DCE, including SNR and image sharpness. This is promising for allowing comprehensive quantitative evaluation of carotid atherosclerosis in a short scan time. Future work will focus on application and validation in patients with carotid atherosclerosis.Acknowledgements
No acknowledgement found.References
1 Kerwin, W. S., Oikawa, M., Yuan, C., Jarvik, G. P. & Hatsukami, T. S. MR imaging of adventitial vasa vasorum in carotid atherosclerosis. Magn Reson Med 59, 507-514, doi:10.1002/mrm.21532 (2008).
2 Yuan, C., Oikawa, M., Miller, Z. & Hatsukami, T. MRI of carotid atherosclerosis. Journal of nuclear cardiology 15, 266-275 (2008).
3 Cai, J.-M. et al. Classification of human carotid atherosclerotic lesions with in vivo multicontrast magnetic resonance imaging. Circulation 106, 1368-1373 (2002).
4 Christodoulou, A.G., Shaw, J.L., Sharif, B., Li, D. A general low-rank tensor framework for high-dimensional cardiac imaging: application to time-resolved T1 mapping. 24th Annual Meeting of ISMRM, Singapore, 2016. Abstract 3032.
5 Xie, Y., Christodoulou, AG., Wang, N., Li, D. Quantitative Multi-Contrast Atherosclerosis Characterization (qMATCH): Comprehensive Quantitative Evaluation of Atherosclerosis in a Single-Scan. 25th Annual Meeting of ISMRM, Honolulu, HI, USA, 2017. Abstract 5148.
6 Wang, N., Christodoulou, AG., Xie, Y., Li, D. Quantitative 3D Dynamic Contrast Enhanced (DCE) Imaging of Carotid Vessel Wall by Fast T1 Mapping. 25th Annual Meeting of ISMRM, Honolulu, HI, USA, 2017. Abstract 7331.
Figures