1312
Reconstruction of motion affected prostate MRSI data using navigators and compressed sensing1Dayananda Sagar Institutions, Bangalore, Karnataka, India, 2Center for Magnetic Resonance Research and Department of Radiology, University of Minnesota, Minneapolis, MN, United States, 3Department of Radiology, Columbia University Medical Centre, New York, NY, United States
Synopsis
This work focuses on reconstruction of 2D prostate in vitro and in vivo MRSI data. Motion affected phase encodes are tracked using a free induction decay navigator. The proposed work utilizes Compressed Sensing (CS) reconstruction technique to compensate for the loss of motion affected information. Comparison between data without motion considered as ground truth (GT) is performed with data with motion and CS reconstructed data. Qualitative and quantitative performance measures indicate improvement in spectral quality with the application of the navigator led CS MRSI reconstruction. Current and future work involves the application of this method on an increased sample size.
Purpose:
To demonstrate robust reconstruction of prostate MRSI data in the presence of navigator sensed motion using Compressed Sensing (CS)1. This work demonstrates the application of MRSI CS reconstruction in a novel scenario where the missing samples in MRSI raw data are dictated by motion rather than driven by acceleration of acquisition alone.Methods:
Navigator based MRSI acquisition:
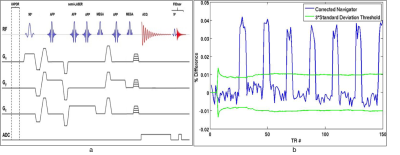
Acquisitions were conducted on whole-body 7 T research MRI system (Siemens, Erlangen, Germany) using a semi-LASER2 sequence (TR/TE = 1900/70ms) with VAPOR3 water suppression and MEGA4 lipid suppression. Free induction decay (FID) navigators5 were employed to track motion during acquisition. Data for each TR was acquired with an FID navigator signal that allowed identification of corrupted points in k-space. Figure 1a shows the pulse sequence diagram with the navigator included while figure 1b shows an example navigator time-course for a scan. During the acquisition, the navigator was corrected for baseline drift throughout the scan due to thermal effects in the gradient coils and a running standard deviation was maintained inclusive of all accepted points. The acquisition was marked as corrupted if the navigator exceeded the threshold and guided the reconstruction to ignore these data points. A ‘sampling mask’ was thus constructed for the reconstruction with 1’s and 0’s representing samples without and with motion corruption. Prospective MRSI data with and without motion was acquired on an in vitro phantom and a volunteer. A custom 18 L phantom (30x45x19cm) was made from cellulose acetate butyrate and plexiglass (Phantom Laboratory, Salem, NY) and fitted with a 9 cm long cylinder running along its length through the center of the phantom with a 3 cm diameter to represent the position and orientation of a human rectum6.
Compressed Sensing Reconstruction:
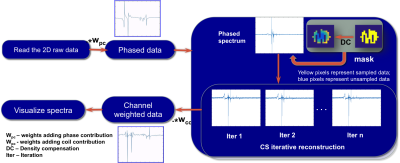
The 16 channel raw data was reconstructed using a motion derived sampling mask. CS based iterative reconstruction using non-linear conjugate gradient was performed exploiting sparsity in one spatial and one temporal dimension using wavelets (Daubechies level 4). Reconstruction was performed with total variation regularization with eight iterations as previously detailed in ref7 . CS reconstructed data was channel weighted to obtain the final reconstructed spectra. The pipeline followed to reconstruct is as shown in the figure 2. The method was demonstrated on data with and without motion on the in vitro phantom data. Fully sampled data without motion was considered as ground truth (GT), while data with motion was reconstructed using CS MRSI. The data with motion and CS reconstruction were compared with GT. Similar experiments were conducted on the in vivo data. RMSE values were calculated for both cases. The code implemented for the reconstruction is available online8.
Results:
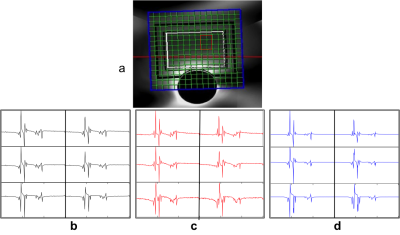
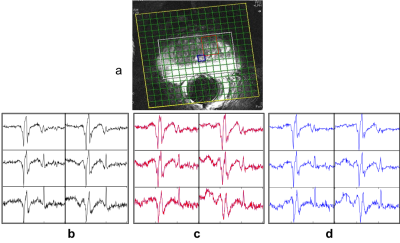
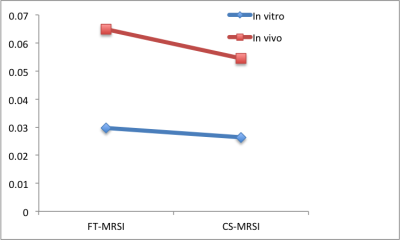
The reconstruction in case of both in vitro and in vivo 2D prostate data was able to reduce the noise in the spectra. The resulting spectra were similar to that of reference spectra of GT data as shown in figure 3 for the case of in vitro phantom data and figure 4 for in vivo experiments. It can be observed that the SNR of the in vitro phantom data is qualitatively better than that of in vivo as expected. This is also reflected in CS reconstructions and RMSE values. The RMSE values for these reconstructions are graphed in figure 5. RMSE observed for the CS MRSI reconstructed data was lesser than that of the data with motion as compared with data without motion. Amplitudes of the metabolites in the region of interest (ROI) (shown in red in figure 3a) are similar to those in the GT case as shown in figures 3(b-d) for the in vitro phantom. Spectra without motion for the voxels in the ROI are plotted in black color, Spectra with motion in red and CS reconstructed spectra in blue as shown in figure 3 (in vitro phantom) and figure 4 (in vivo).Discussion and Conclusion:
Navigator led motion correction using CS MRSI method has been demonstrated on in vitro phantom and an in vivo data set. The qualitative and quantitative evaluations validate the improvement in motion mitigation using this approach. Future work involves applying this method on more data sets to gain statistical significance and extension to three dimensional MRSI.Acknowledgements
This work was supported by Department of Science and Technology (DST), Govt. of India under the program Technology Systems Development (TSD) for the project “Novel acquisition and reconstruction strategies to accelerate magnetic resonance imaging using compressed sensing”, No: DST/TSG/NTS/2013/100-G; NCI R01 CA155268, NIBIB P41 EB015894References
[1] Lustig M, Donoho D, Pauly JM, “Sparse MRI: The application of compressed sensing for rapid MR imaging”, Magnetic Resonance in Medicine 2007; 58(6):1182-1195
[2] Scheenen, Tom WJ, Arend Heerschap, and Dennis WJ Klomp. "Towards 1H-MRSI of the human brain at 7T with slice-selective adiabatic refocusing pulses." Magnetic Resonance Materials in Physics, Biology and Medicine 21.1-2 (2008): 95.
[3] Tkáč, I., et al. "In vivo 1H NMR spectroscopy of rat brain at 1 ms echo time." Magnetic resonance in medicine 41.EPFL-ARTICLE-177519 (1999): 649-656.
[4] Mescher, M., et al. "Simultaneous in vivo spectral editing and water suppression." NMR in Biomedicine 11.EPFL-ARTICLE-177509 (1998): 266-272.
[5] Kober, Tobias, et al. "Head motion detection using FID navigators." Magnetic resonance in medicine 66.1 (2011): 135-143.
[6] Ertürk, M. Arcan, et al. "Development and evaluation of a multichannel endorectal RF coil for prostate MRI at 7T in combination with an external surface array." Journal of Magnetic Resonance Imaging 43.6 (2016): 1279-1287.
[7] Geethanath, Sairam, et al. "Compressive sensing could accelerate 1H MR metabolic imaging in the clinic." Radiology262.3 (2012): 985-994.
[8] https://github.com/mirc-dsi/IMRI-MIRC/tree/master/CSMRSI_recon
Figures