0430
Simultaneous detection and identification of MR artifact types in whole-body imaging1Section on Experimental Radiology, University Hospital of Tuebingen, Tuebingen, Germany, 2Institute of Signal Processing and System Theory, University of Stuttgart, Stuttgart, Germany, 3Department of Radiology, University Hospital of Tuebingen, Tuebingen, Germany
Synopsis
Varying acquisition and reconstruction conditions as well as long examination times make MRI susceptible to various kinds of artifacts. If suitable correction techniques are not available/applicable, if human experts who judge the achieved quality are not present or for epidemiological cohort studies in which a manual quality analysis of the large database is impracticable, an automated detection and identification of these artifacts is of interest. Convolutional neural networks with residual and inception layers localize and identify occurring artifacts. Artifacts (motion and field inhomogeneity) can be precisely identified with an accuracy of 92% in a whole-body setting with varying contrasts.
Introduction
Depending on the chosen sequence type and contrast weighting, MRI is more or less susceptible to several types of image artifacts. In order to guarantee high data quality, arising artifacts need to be detected as early as possible to seize appropriate countermeasures. Due to the manifold of possible occurring artifacts not all precautions can be considered, hence leaving a chance of artifacts being present in the final image. It is the task of a human MR specialist to appreciate the level of achieved image quality with respect to the underlying application. This analysis can be a time-demanding and cost-intensive process. Insufficient image quality may demand an additional examination decreasing patient comfort and throughput. Thus, a prospective quality assurance is highly desired.
In the context of large epidemiological cohort studies such as UK Biobank1 or German National Cohort2, reliable image quality has to be guaranteed. However, the amount and complexity exceed practicability for a manual analysis. Thus, a retrospective quality assessment/control is desired.
An automated and reference-free quality analysis is preferred. Previously proposed approaches for automated image quality analysis required the existence of a reference image or were only focused on specific scenarios3-5. Reference-free approaches6-10 are mainly metric-based driven to evaluate quality on a coarse level.
In a previous work, we showed the potential of a deep learning network to perform the task of automatic reference-free motion artifact detection11,12. In this work, we extend this concept to a multi-class scenario for identification of motion and magnetic field inhomogeneity artifacts in a whole-body scenario and in images exhibiting different contrast weightings. We propose two convolutional neural network architectures for this task and investigate their performance.
Material and Methods
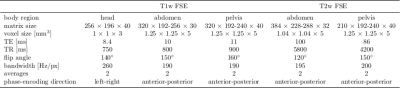
MR images were acquired on a 3T PET/MR (Biograph mMR, Siemens) from 18 healthy volunteers (3 female, 25±8 years) with a T1w and T2w FSE sequence. The acquisition parameters for the respective body regions (head, abdomen, pelvis) are depicted in Tab.1. In each body region and contrast two acquisitions were performed: a reference and a motion-corrupted (head, hips movement and breathing) scan. For the T2w sequence, magnetic field inhomogeneity artifacts were acquired with manually disturbed B0 shimming. All images are normalized into an intensity range of 0 to 1 and partitioned into 50% overlapping patches of size 80x80 (APxLR).
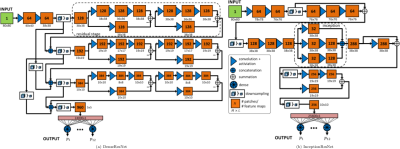
The proposed convolutional neural network (CNN) architectures are depicted in Fig.2. The DenseResNet is inspired by combining the ideas of DenseNet13 and ResNet14. It consists of five stages. A first convolutional stage, followed by three residual stages and a final fully connected output stage. The dense connections act as a bypass of feature maps from previous layers to deep layers enabling a joint estimation on coarse- and fine-grained feature maps in deeper layers. Residual shortcuts feed-forward feature maps to provide residual mapping and to enable ensemble learning. The residual blocks are built up of a 1x1 convolutional layer wrapped around the 3x3 convolutional layer acting as a bottleneck structure to merge feature maps by subtraction across channels.
The InceptionResNet is inspired by the GoogLeNet15 which uses inception layers. The idea is to cover larger spatial areas by multi-scale convolutions and keeping smaller image portions for deeper levels. The architecture consists of three convolutional stages with inception modules on the second and third stage. In the first and second stage an additional residual path forwards the feature map to a deeper level.
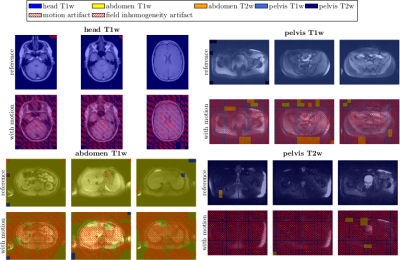
Both architectures are trained by a leave-one-subject-out cross-validation to output probability values $$$p_i$$$ for each of the twelve classes (see Fig.3). Categorical cross-entropy is minimized for given learning rate, $$$\ell_2$$$ regularization and dropout. Parameter ranges are estimated by the Baum-Haussler rule16 with a grid-search optimization. Testing was performed on the left-out subject to investigate accuracy and confusion matrices.
Results and Discussion
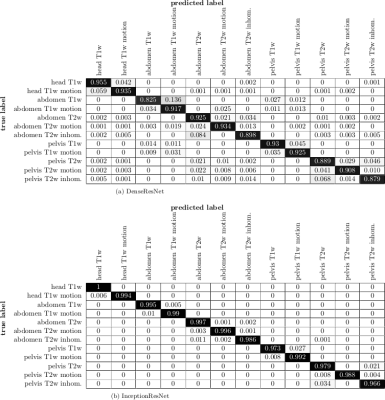
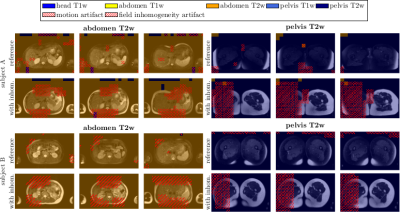
Fig.3 depicts the output probability maps in three exemplary slices of one subject for DenseResNet. Fig.4 compares the performance in InceptionResNet amongst subjects. The respective body region and contrast weighting (color) as well as artifact (grid) are always correctly identified in each foreground patch with some minor false-positives in the background and between abdomen and pelvis. This observation is also supported by the block-diagonal structure of the confusion matrices (Fig.5). Average accuracies over cross-validations of 90.8±6.1% and 91.9±9.6% were achieved by the DenseResNet and InceptionResNet, respectively, with fluctuations according to degree and manifestation of artifacts amongst subjects. Both architectures provided nearly similar performance.Conclusion
A CNN can be trained to automatically identify different types of artifacts in a whole-body scenario with changing contrast weighting achieving high accuracies of >90%. This is of interest for automatic MR quality controls.Acknowledgements
No acknowledgement found.References
[1] W. Ollier, T. Sprosen, and T. Peakman, “UK Biobank: from concept to reality,” Pharmacogenomics, vol. 6, no. 6, pp. 639-646, 2005. [2] F. Bamberg, H. U. Kauczor, S. Weckbach, C. L. Schlett, M. Forsting, S. C. Ladd, K. H. Greiser, M. A. Weber, J. Schulz-Menger, T. Niendorf, T. Pischon, S. Caspers, K. Amunts, K. Berger, R. Bulow, N. Hosten, K. Hegenscheid, T. Kroncke, J. Linseisen, M. Gunther, J. G. Hirsch, A. Kohn, T. Hendel, H. E. Wichmann, B. Schmidt, K. H. Jockel, W. Hoffmann, R. Kaaks, M. F. Reiser, and H. Volzke, “Whole-Body MR Imaging in the German National Cohort: Rationale, Design, and Technical Background,” Radiology, vol. 277, no. 1, pp. 206–20, 2015. [3] J. Oh, S. I. Woolley, T. N. Arvanitis, and J. N. Townend, “A multistage perceptual quality assessment for compressed digital angiogram images,” IEEE Trans. Med. Imaging., vol. 20, no. 12, pp. 1352–61, 2001. [4] J. Miao, D. Huo, and D. L. Wilson, “Quantitative image quality evaluation of MR images using perceptual difference models,” Med. Phys., vol. 35, no. 6, pp. 2541–53, 2008. [5] A. Ouled Zaid and B. B. Fradj, “Coronary angiogram video compression for remote browsing and archiving applications,” Comput. Med. Imaging. Graph., vol. 34, no. 8, pp. 632–41, 2010. [6] B. Mortamet, M. A. Bernstein, Jr. Jack, C. R., J. L. Gunter, C. Ward, P. J. Britson, R. Meuli, J. P. Thiran, and G. Krueger, “Automatic quality assessment in structural brain magnetic resonance imaging,” Magn. Reson. Med., vol. 62, no. 2, pp. 365–72, 2009. [7] J. P. Woodard and M. P. Carley-Spencer, “No-reference image quality metrics for structural MRI,” Neuroinformatics, vol. 4, no. 3, pp. 243–62, 2006. [8] M. D. Tisdall and M. S. Atkins, “Using human and model performance to compare MRI reconstructions,” IEEE Trans. Med. Imaging., vol. 25, no. 11, pp. 1510–7, 2006. [9] D. Atkinson, D. L. Hill, P. N. Stoyle, P. E. Summers, and S. F. Keevil, “Automatic correction of motion artifacts in magnetic resonance images using an entropy focus criterion,” IEEE Trans. Med. Imaging, vol. 16, no. 6, pp. 903–10, 1997. [10] K. P. McGee, A. Manduca, J. P. Felmlee, S. J. Riederer, and R. L. Ehman, “Image metric-based correction (autocorrection) of motion effects: analysis of image metrics,” J. Magn. Reson. Imaging, vol. 11, no. 2, pp. 174–181, 2000. [11] T. Küstner, A. Liebgott, L. Mauch, P. Martirosian, F. Bamberg, K. Nikolaou, B. Yang, F. Schick, and S. Gatidis, “Automated reference-free detection of motion artifacts in magnetic resonance images,” Magn. Reson. Mater. Phys., Biol. Med., Sep 2017. [12] T. Küstner, A. Liebgott, L. Mauch, P. Martirosian, K. Nikolaou, F. Schick, B. Yang, and S. Gatidis, „Automatic reference-free detection and quantification of MR image artifacts in human examinations due to motion,” ISMRM Proceedings, p. 1278, 2017. [13] G. Huang, Z. Liu and K. Q. Weinberger, “Densely connected convolutional networks,” CoRR, vol. abs/1608.06993, 2016. [14] K. He, X. Zhang, S. Ren and J. Sun, “Deep residual learning for image recognition,” CoRR, vol. abs/1512.03385, 2015. [15] C. Szegedy, V. Vanhoucke, S. Ioffe, J. Shlens and Z. Wojna, “Rethinking the inception architecture for computer vision,” CoRR, vol. abs/1512.00567, 2015. [16] E.B. Baum and D. Haussler, “What Size Net Gives Valid Generalization?,” Neural Comput., vol. 1, no. 1, pp. 151–160, Mar. 1989.Figures