3631
A Deconvolution Method for Improved CBF Quantification in 3D-GRASE ASL1Biomedical Engineering, Stony Brook University, Stony Brook, NY, United States, 2Radiology, Stony Brook University, Stony Brook, NY, United States
Synopsis
Pseudo-continuous arterial spin labeling (pCASL) with segmented 3D-GRASE acquisition is widely accepted as the optimal ASL technique. However, the method suffers from blurring along the partition direction caused by point spread function (PSF) broadening. In this study, a PSF deconvolution method for pCASL images with 3D-GRASE acquisition is developed and evaluated in simulations and in-vivo experiments. The deconvolution method greatly reduces the effects of the PSF and recover the perfusion signal for segmentation factors of at least 2PAR x 2PE. The proposed deconvolution method improves the accuracy of cerebral blood flow quantification and facilitates the use of lower segmentation factors.
Purpose:
Arterial spin labeling (ASL) MRI is a noninvasive technique to measure cerebral blood flow (CBF)1. The 2015 ASL white paper designated pseudo-continuous ASL (pCASL) with segmented 3D-GRASE to be the optimal ASL technique2. However, 3D-GRASE acquisition suffers from blurring along the partition (PAR) direction caused by point spread function (PSF) broadening from long echo trains. High segmentation factors along phase-encoding (PE) and PAR directions reduce blurring but also reduce temporal resolution, which may increase sensitivity to subject motion3. Additionally, 3D-GRASE ASL produces lower CBF values than conventional 2D approach potentially due to pseudo-diffusion and PSF. In this study, a post-processing deconvolution method is proposed and evaluated, which improves the accuracy of CBF quantification.Methods:
The effect of image blurring along the PAR direction in 3D-GRASE images can be modeled by convolution of the true image with the PSF. Non-blind deconvolution was applied iteratively according to,
$$D^{i+1}=\min\left[\parallel P\ast D^{i}-B\parallel_2^2+\lambda^{2}\parallel D^{i}-R \parallel_2^2+\lambda^{2}\parallel w\left(\nabla D^{i}\right)\parallel_2^2\right]\qquad.$$
Where D is the deblurred image, P is the point spread function, B is the measured image, λ is the regularization parameter determined by the L-curve method, R is a local smoothness constraint, and w is related to structural gradient to preserve anatomical integrity.
PSF’s for segmentation factors of 1PAR x 2PE, 2PAR x 2PE, 3PAR x 2PE, and 4PAR x 2PE were estimated using extended phase graphs4. Pseudo-diffusion effect from intravascular tagged water signal (~30% 5) was accounted for by augmented EPG6.
A numerical perfusion phantom was generated based on high resolution brain MPRAGE images with CBF values of 60 and 20 ml/100g/min assigned for GM and WM respectively. The deconvolution method was assessed through simulations at different levels of signal to noise ratio (SNR). Dispersion was calculated as the standard deviation of CBF in GM and estimation bias was calculated as,
$$Bias=\left[\overline{CBF}_{GM}^{True}-\overline{CBF}_{GM}^{Est}\right]\div \overline{CBF}_{GM}^{True}\times100\% \qquad.$$
Six subjects were recruited for this IRB approved study. All experiments were performed on 3T Siemens scanners. High resolution MPRAGE images with voxel size of 1x1x1 mm3 were acquired for brain structure. A segmentation factor of 2 on PE was adopted for all 3D-GRASE images. Segmentation factors on PAR direction were 1, 2, 3, and 4. Other MR parameters were: TR of 4 seconds; 80x64x40 matrix; 3 mm isotropic resolution; RF flip angle of 120°; labeling time of 1600 ms; post-labeling delay of 1500 ms. Reference images were acquired for each segmentation factor with a TR of 8 seconds. The total acquisition times were kept at 6 minutes.
ASL images were realigned, coreregistered, and smoothed using ASLtbx7. GM, WM, and CSF probability maps were generated using SPM12. Deconvolution was applied on the difference image using PSFs with and without pseudo-diffusion effect.
Results & Discussions:
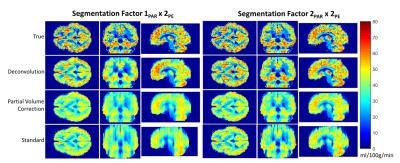
Fig. 1 displays the CBF maps from simulations. Deconvolution nearly eliminated the blurring artifact and recovered the perfusion map for segmentation factors of 2PAR x 2PE and greater. For segmentation factor of 1PAR x 2PE, deconvolution only partially resolved the blurring. The bias and dispersion for different segmentation factors and SNR levels is shown in Fig. 2. Deconvolution reduced CBF bias for all segmentation factors, and brings it within the physiological noise level of ASL studies except for 1PAR x 2PE segmentation. As expected, bias showed little dependency on SNR and the deconvolution slightly increases the dispersion of CBF quantification.
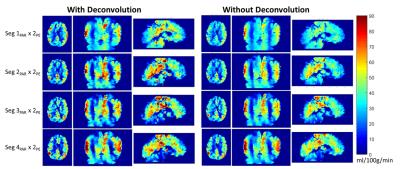
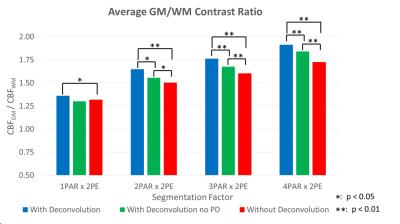
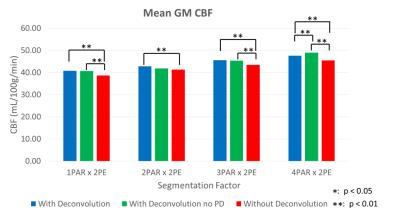
Fig. 3 displays the typical result from a volunteer subject. The proposed method reduces the blurring along the PAR direction and significantly improves the GM/WM contrast ratio (Fig. 4). These results indicate that for segmentation factors of at least 2PAR x 2PE, the proposed deconvolution method is capable of reconstituting the perfusion signal. This is further supported by the mean GM CBF estimated across all six subjects (Fig. 5). Comparing deconvolution using PSF with and without pseudo-diffusion, the underestimation of GM CBF is still significant until very high segmentation factors are adopted. This is consistent with the notion that PSF with pseudo-diffusion is much broader than conventional PSF due to T1/T2 relaxation. Meanwhile, deconvolution with the pseudo-diffusion effect further improves the GM/WM contrast ratio as demonstrated in Fig 4.
Conclusion:
The proposed PSF deconvolution method improves CBF quantification based on pCASL with segmented 3D-GRASE acquisition. The performance of the method was evaluated in simulation and in-vivo experiment. This study demonstrated that PSF deconvolution can recover the CBF maps for segmentation factors of 2PAR x 2PE and greater, while significantly reducing the bias of quantification. Studies employing pCASL with segmented 3D-GRASE acquisition would greatly benefit from the proposed method.Acknowledgements
No acknowledgement found.References
1. Detre, John A., et al. "Applications of arterial spin labeled MRI in the brain." Journal of Magnetic Resonance Imaging 35.5 (2012): 1026-1037.
2. Alsop, David C., et al. "Recommended implementation of arterial spin-labeled perfusion MRI for clinical applications: A consensus of the ISMRM perfusion study group and the European consortium for ASL in dementia." Magnetic resonance in medicine 73.1 (2015): 102-116.
3. Vidorreta, Marta, et al. "Evaluation of segmented 3D acquisition schemes for whole-brain high-resolution arterial spin labeling at 3 T." NMR in Biomedicine27.11 (2014): 1387-1396.
4. Weigel, Matthias. "Extended phase graphs: Dephasing, RF pulses, and echoes-pure and simple." Journal of Magnetic Resonance Imaging 41.2 (2015): 266-295.
5. St Lawrence, Keith S., et al. "A two-stage approach for measuring vascular water exchange and arterial transit time by diffusion-weighted perfusion MRI." Magnetic resonance in medicine 67.5 (2012): 1275-1284.
6. He, Xiang, et al. Diffusion Sensitivity of ASL Perfusion with 3D-GRASE Readout. International Society of Magnetic Resonance in Medicine 23rd Annual Meeting, (2015): Toronto.
7. Wang, Ze, et al. "Empirical optimization of ASL data analysis using an ASL data processing toolbox: ASLtbx." Magnetic resonance imaging 26.2 (2008): 261-269.
Figures