3540
TORTOISE v3: Improvements and New Features of the NIH Diffusion MRI Processing Pipeline1Quantitative Medical Imaging Section, NIBIB/NIH, Bethesda, MD, United States, 2SQUITS/NICHD/NIH, Bethesda, MD, United States, 3Henry Jackson Foundation, Bethesda, MD, United States
Synopsis
Here we present a series of improvements and new features of the TORTOISE diffusion MRI data processing software (www.tortoisedt.org). TORTOISEv3 has been programmed in C++ and it is now significantly faster, can be batched and it fully benefits from modern multi-core CPU architectures. The DIFFPREP module brings a multitude of new and state-of-art features including DWI denoising, Gibbs ringing removal, and the ability to perform motion and eddy currents distortion correction for very high b-value data. The new DIFFCALC module can perform MAP-MRI propagator estimation and the output can be easily imported in other software packages for statistical analysis and atlas creation.
Introduction
Diffusion MRI modalities, including diffusion tensor imaging (DTI), are commonly used for both research and diagnostic purposes. Diffusion MRI data suffer from a number of artifacts, such as bulk subject motion, eddy current distortion, and susceptibility induced EPI distortion [1-2]. Appropriate pre-processing of diffusion weighted images and robust processing of diffusion tensor data is vital for accurate quantitative analysis. The TORTOISE (Tolerably Obsessive Registration and Tensor Optimization Indolent Software Ensemble) software [3], is an integrated set of diffusion MRI processing tools developed by NIH researchers and has been publicly available for about a decade (www.tortoisedti.org). The original TORTOISE was mostly written in the IDL programming language and was optimized for diffusion tensor data. As DWI acquisition and diffusion modeling has advanced significantly, novel and improved strategies to perform corrections and process DWI data are needed. The new TORTOISE that we present here has been completely rebuilt to take advantage of improved computational approaches and added new features.Methods and Results
Table 1 summarizes the new features of TORTOISEv3. Among these new features, the remainder of this manuscript will focus on DIFFPREP’s new strategies to perform high b-value motion & eddy currents distortion correction and its MAP-MRI features.
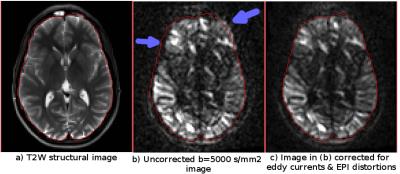
High b-value motion & eddy currents distortion correction: DIFFPREP’s strategy for low-shell motion & eddy currents distortion is to register the DWIs to a b=0 s/mm2 image with a mutual information metric and a parsimonious quadratic/cubic eddy currents transformation model [1]. This strategy however fails for high b-value data, where the contrast between DWIs and the b=0 s/mm2 image are significantly different. FSL’s Eddy tool [8] overcomes this issue by using the entire diffusion set to make predictions of how the image contrast should be. Even though this approach has been proven to be superior to the previous eddy_correct [9], its performance is expected to depend on the DWI acquisition scheme. Specifically it could not perform properly if the number of images is low and/or the diffusion directions are not nicely distributed. The new DIFFPREP is designed to be less dependent on experimental design. The low b-value data (0-1,200 s/mm2), which obey the tensor model and have high SNR, are corrected with the approach described in [1] which has proven to be very robust in this regime. For mid level b-values (e.g. 2,500 s/mm2), the previously corrected low shell volumes are used to estimate the corresponding diffusion tensors and for a given mid b-value and gradient direction, the corresponding DWI is synthesized from the tensor model and the actual image is registered to its synthesized version. For high b-value data, all the previously corrected images are used to perform a MAP-MRI [7] based propagator estimation and another image is synthesized with the same q as the current DWI. The registration from actual DWI to MAP-MRI synthesized version is initialized differently: The motion components are initialized using a moving-window linear regression. The eddy currents correction parameters are initialized using the entire corrected DWI set with an approach similar to Eddy’s [8] second level modeling by performing a linear regression on gradient strength and x, y, z, components of the diffusion gradients as inputs. DIFFPREP’s new strategy does not necessitate blip-up blip-down or full diffusion sphere acquisitions, is robust and yields very high quality corrections. However, it is not designed for single-shell high b-value HARDI acquisitions. An example correction is displayed in Figure 1.
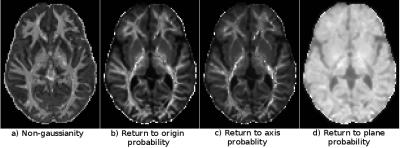
MAP-MRI features: MAP-MRI [7] is a diffusion propagator estimation methodology, which uses an analytical, orthonormal basis set to describe the probabilities for the positions of water molecules. With this strategy, once the diffusion propagator is estimated, it is straightforward to derive several maps/stains about the tissue microstructure. Non-gaussianity map, which describes the deviations from a gaussian diffusion described by the tensor model, the propagator anisotropy, which describes deviations from an isotropic propagator and return to the origin probability, which describes the probability that a particle will return to its starting position and which is directly inversely proportional to pore size are examples of such maps. Figure 2 displays a non-gaussianity map, return to origin, return to axis and return to plane maps.
Conclusions
The new TORTOISE, which can be downloaded from www.tortoisedti.org, is completely rebuilt with a new architecture and significantly improves upon its older counterpart by enhancing its existing features and adding several additional ones to accommodate the current processing needs of clinicians and diffusion MRI research scientists.Acknowledgements
Support for this work included funding from the Congressionally Directed Medical Research Programs (CDMRP).References
1. Rohde, G.K., Barnett, A.S., Basser, P.J., Marenco, S., and Pierpaoli, C. (2004) Comprehensive approach for correction of motion and distortion in diffusion-weighted MRI. Magn Reson Med. 51:103-114.
2. Wu, M., Chang, L.-C., Walker, L., Lemaitre, H., Barnett, A.S., Marenco, S., and Pierpaoli, C. (2008) Comparison of EPI Distortion Correction Methods in Diffusion Tensor MRI Using a Novel Framework. MICCAI 2008, Part II, LNCS 5242, pp. 321-329.
3. C. Pierpaoli, L. Walker, M. O. Irfanoglu, A. Barnett, P. Basser, L-C. Chang, C. Koay, S. Pajevic, G. Rohde, J. Sarlls, and M. Wu. (2010) TORTOISE: an integrated software package for processing of diffusion MRI data. In ISMRM 18th Annual Meeting and Exhibition, Stockholm, Sweden. May 1-7, 2010.
4. Veraart, J.; Novikov, D.S.; Christiaens, D.; Ades-aron, B.; Sijbers, J. & Fieremans, E. Denoising of diffusion MRI using random matrix theory. NeuroImage, 2016, , doi: 10.1016/j.neuroimage.2016.08.016.
5. Kellner, E., Dhital, B., Kiselev, V. G. and Reisert, M. (2016), Gibbs-ringing artifact removal based on local subvoxel-shifts. Magn. Reson. Med., 76: 1574–1581.
6. Irfanoglu MO, Modi P, Nayak A, Hutchinson EB, Sarlls J, and Pierpaoli C. DR-BUDDI (Diffeomorphic Registration for Blip-Up blip-Down Diffusion Imaging) method for correcting echo planar imaging distortions. NeuroImage 106 (2015) 284–299.
7. Özarslan E, Koay CG, Shepherd TM, Komlosh ME, Irfanoglu MO, Pierpaoli C, and Basser PJ. (2013) Mean Apparent Propagator (MAP) MRI: a novel diffusion imaging method for mapping tissue microstructure. Neuroimage 78:16-32. 8.
8. Jesper L. R. Andersson and Stamatios N. Sotiropoulos. An integrated approach to correction for off-resonance effects and subject movement in diffusion MR imaging. NeuroImage, 125:1063-1078, 2016.
9. Mark S. Graham, Ivana Drobnjak and Hui Zhang. Realistic simulation of artefacts in diffusion MRI for validating post-processing correction techniques. NeuroImage, 125:1079-1094, 2015.
Figures