1942
An optimised 2D MPRAGE sequence for T1 contrast in the fetal brain: application to slice to volume reconstruction and multiband acceleration1Division of Imaging Sciences & Biomedical Engineering, King's College London, London, United Kingdom, 2Translational and Molecular Imaging Institute, Icahn School of Medicine at Mount Sinai, New York, United States
Synopsis
Ultrafast single-shot T2 weighted images are common practise in fetal MR exams. However, there is limited experience with fetal T1 acquisitions.
In this study, a 2D gradient echo sequence with an adiabatic inversion was optimized to be robust to fetal motion and to preserve contrast. We also explore slice to volume registration and super resolution reconstruction methods to enhance the resolution, and we show pilot data from a multiband accelerated version of the above-mentioned sequence.
Introduction
There is growing demand for in utero brain imaging for diagnosis and research. The workhorse sequence is single-shot-fast-spin-echo (ssFSE), predominantly for T2w imaging, but also applied with inversion recovery preparation for T1w imaging1. However, these T1w sequences are not as sensitive to pathology as spoiled gradient-echo (GE), and as currently performed the latter is very motion sensitive and often requires a breath-hold. Yet a combination of T1w and T2w imaging is required for secure and robust diagnosis.
In this study, we optimize a 2D single-shot MPRAGE sequence to image the fetal brain seeking high contrast and robustness to motion at the same time. We also explore different options for multiband (MB) imaging2, and use slice-to-volume reconstruction (SVR)3 to achieve full 3D anatomical depiction.
Methods
All data was acquired using a Philips Achieva 3T system with a 32 channel cardiac array coil.
In our approach a 2D single-shot slice is acquired to freeze fetal motion, whereas a non-selective adiabatic inversion is used for magnetization preparation to achieve robustness to motion during the inversion time (TI). The approach differs from previous work1, where a short TI in combination with slice selective excitation was used to contain motion.
The application of a train of global inversions at intervals TS sets up a dynamic steady state in which each tissue follows its own T1 evolution. This steady state is sampled one slice at a time which disturbs the magnetisation, and then the steady state is reasserted during subsequent TS periods. An inversion time delay of TI seconds is introduced between the adiabatic pulse and the start of the gradient acquisition readout consisting of a train of pulses of flip angle α, separated by a time TR. We found that it was not beneficial to introduce a post-readout delay, so for a fixed number of phase encodes (which in our study was approximately 180, often halved to 90 because of SENSE 2), TS is predetermined by TI and TR.
Simulations were performed using literature T1 values4 for CSF and neonatal white and grey matter at 3T, and a search of parameter space conducted to seek optimal settings. Figure 1 shows grey-white matter contrast (a) and individual tissue signals (b) obtained for α = 15o and TR = 10 ms using SENSE 2. Contrast is maximised for an inversion time of 3.5 seconds. Figure 1b shows the evolution of the signal as a function of the readout duration. Although centric ordering would be expected to enhance contrast, we found that linear ordering produces similar results (see Figure 3b, details in caption), provided the readout duration was around two seconds or less. In all cases minimum bandwidth readout was used to maximise SNR. After some testing a choice for TI of 2.5 sec was adopted, which represents a good compromise between contrast and acquisition efficiency.
Stacks of T1 weighted data were acquired transverse, sagittal and coronal to the fetal head, all with in-plane resolution of 1.5x2mm2 and slice thickness of 5 mm. All data was then reconstructed jointly at 0.75 mm isotropic using SVR. To speed up acquisition, multiband with the appropriate shift pattern5 was implemented and applied to the GE module and reconstruction was performed using CG-SENSE6. Finally, we extended the SVR framework3 for multiband acquisitions to ensure that slices belonging to the same excitation are registered together.
The approach was tested on 15 fetuses ranging from 24 to 40 weeks gestational age (wGA). Written informed consent was obtained from all participants.
Results
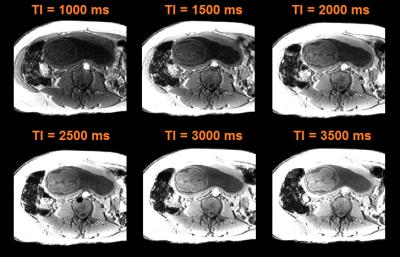
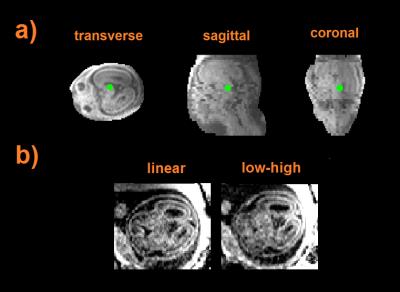
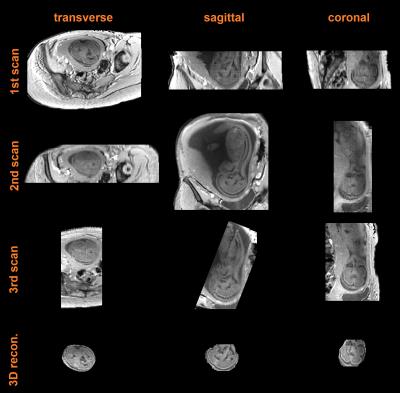
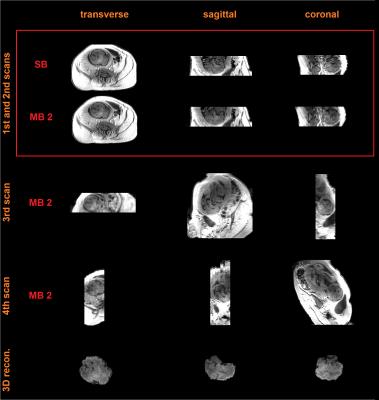
Figure 2 shows data from a fetus 25 wGA acquired at increasing inversion times providing evidence that good contrast exists at 2.5 sec. Figure 3a shows data from the same fetus as in Figure 2 (TI = 3.5 sec) where, in spite of substantial inter slice motion (visible mostly in the sagittal view), contrast is preserved in the native transverse plane. Figure 4 shows single band (SB) data from a fetus of 25 wGA acquired transverse (first row), sagittal (second) and coronal (third) to the fetal head. The fourth row shows SVR reconstruction. Figure 5 shows SB (first row) versus MB2 data (second, third and fourth rows, details in caption). Image quality is preserved, despite a reduction of scan time from 84 sec to 42 sec. The fifth row shows the SVR reconstruction.Discussion and Conclusion
In this study we have optimized a fetal T1 single-shot 2D MPRAGE sequence with non-selective inversion that is robust to motion and suitable for SVR reconstruction. We are exploring different options for multiband in combination with SENSE and Partial Fourier.Acknowledgements
We would like to acknowledge the following funding sources: MRC strategic grant (MR/K006355/1), GSTT Biomedical Research Centre and European Research Council funded dHCP project.References
1 Malamateniou C, McGuinness A K, Allsop J M et al. An Optimized Single-Shot T1 weighted Inversion-Recovery Sequence for Improved Fetal Brain Anatomy Delineation. Radiology. 2011;258(1):229-235.
2. Larkman D J, Hajnal J V, Herlihy A H et al. Use of multicoil arrays for separation of signal from multiple slices simultaneously excited. Journal of Magnetic Resonance Imaging. 2001;13(2):313-317.
3. Kuklisova-Murgasova M, Quaghebeur G, Rutherford M A et al. Reconstruction of fetal brain MRI with intensity matching and complete outlier removal. Medical Imaging Analysis. 2012;16(8):1550-1564.
4. Williams L, Gelman N, Picot P A, et al. Neonatal Brain: Regional Variability of in Vivo MR Imaging Relaxation Rates at 3.0T – Initial Experience. Radiology. 2005;235(2):595-603.
5. Breuer F A, Blaimer M, Heidemann R M et al. Controlled aliasing in parallel imaging results in higher acceleration (CAIPIRINHA) for multi-slice imaging. Magnetic Resonance in Medicine. 2005;53(3):684-691.
6. Pruessmann K P, Weiger M, Börnert P et al. Advances in sensitivity encoding with arbitrary k-space trajectories. Magnetic Resonance in Medicine. 2001;46(4):638-651.
Figures