0786
Diffusion MRI of the entire postmortem human spinal cord at microscopic resolution1Duke University, Durham, NC, United States
Synopsis
Diffusion MR imaging of the human spinal cord has become increasingly important in both clinical diagnostics, and research science.1 As MRI methods improve, there is a need to understand the limits of diffusion MRI in the human spinal cord. Here, we present a microscopic resolution diffusion MRI dataset of the entire postmortem human spinal cord, generated from a multi-segment acquisition, using an automated image-processing pipeline. These data provide unique insights for spinal cord research, future diagnostic imaging applications, and for postmortem pathologic evaluation of spinal cord specimens.
Purpose
To investigate the use of microscopic resolution diffusion MR imaging in the whole human spinal cord and explore its potential applications for both clinical diagnostics and scientific research.Methods
An entire human spinal cord (pyramidal decussation to cauda equina) was obtained from an adult male at autopsy. The cord was immersion fixed in 10% formalin, then rehydrated in PBS doped with 2.5 mM gadoteridol following previously published methods.2 Both anatomic and diffusion-weighted MR images of the entire cord were acquired in 7 separate overlapping segments using a 7T Agilent small animal MRI system. Between each acquisition, the cord was advanced through the magnet bore using a custom fabricated gantry insert with precisely machined locking distance marks. Images were reconstructed, corrected for gradient non-linearity, and digitally combined using a custom-developed, fully-automated image-processing pipeline. Diffusion data processing and tractography were performed in DSI Studio.3Results
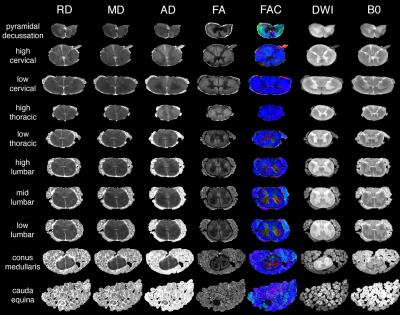
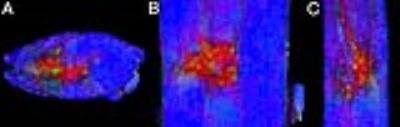
The final composite dataset consists of 50-μm isotropic resolution T2*-weighted anatomic images, and 100-μm isotropic resolution 30-direction diffusion images (b=4000 s/mm2) over a 2 x 2 x 47 cm FOV (Fig. 1). Standard Diffusion Tensor Imaging (DTI)-derived scalar images were generated (Fig. 2), and data were processed for Generalized Q-sampling Imaging (GQI) multi-fiber tractography (Fig. 3-4).3 These data reveal clinically important anatomy not commonly seen on conventional spinal cord MRI, including dorsal and ventral spinal nerve roots (e.g. Fig. 3), individual ventral horn motor nuclei (e.g. Fig 1, d), the central canal (Fig. 1, a), and the anterior white commissure (Fig. 1, e and f). The data also include a spinal cord lesion with signal characteristics typical of subacute to chronic spinal cord injury with associated gliosis (Fig. 5).Discussion
These data provide a unique opportunity to explore the diffusion MRI characteristics of the whole human spinal cord at microscopic resolution. The importance of these data are 2-fold: 1) they provide insight into the upper limits of what may one day be available for diagnostic MRI of the spinal cord, and 2) they demonstrate the level of anatomic detail that can be identified in spinal cord specimens for research and postmortem pathologic assessment. Current applications of this work include: 1) mapping of fiber anatomy for neurosurgical procedures such as dorsal root rhizotomy, 4 2) modeling of fiber activation/inhibition for neuro-stimulators and other neuroprostheses, 5 and 3) pathologic evaluation of spinal cord specimens at autopsy.6Conclusion
We present a postmortem diffusion MRI dataset of the entire human spinal cord at microscopic resolution, made possible by multi-segment acquisition and automated image composition. The data and methods presented here have applications for clinical and pathologic diagnostics, as well as research.Acknowledgements
Funding was provided by NIH/NIBIB grant P41 EB015897 and 1S10OD010683-1.References
1. Naidich TP, Castillo M, Cha S, et al. Imaging of the Spine. Elsevier Health Sciences, Philadelphia, PA; 2011.
2. Johnson GA, Cofer GP, Gewalt SL, Hedlund LW. Morphologic phenotyping with MR microscopy: The visible mouse. Radiology 2002;222(3):789-793.
3. Yeh F-C, Wedeen VJ, Tseng, WY. Generalized q-sampling imaging. IEEE Trans Med Imaging. 2010;29(9):1626-1635.
4. Rahimpour S, Lad SP. Surgical options for atypical facial pain syndromes. Neurosurg Clin N Am. 2016;27(3):365-370.
5. Calabrese E, Hickey P, Hulette C, et al. Postmortem diffusion MRI of the human brainstem and thalamus for deep brain stimulator electrode localization. Hum Brain Mapp. 2015;36(8):3167-3178.
6. Gilmore CP, Geurts JJ, Evangelou N, et al. Spinal cord grey matter lesions in multiple sclerosis detected by post-mortem high field MR imaging. Mult Scler. 2009; 15(2):180-188.
Figures